GMOD
News/JBrowse2 v1.1.0 Released
Contents
- 1 We’re pleased to announce a new release of JBrowse Web!
We’re pleased to announce a new release of JBrowse Web!
(Reposted by permission from https://jbrowse.org/jb2/blog/2021/03/29/v1.1.0-release
Changed callbacks language from JavaScript to Jexl
To allow users to safely and seamlessly share advanced configurations in sessions, we now use Jexl to express configuration callbacks. Note that this is a breaking change, function()-style callbacks will no longer work.
For details, see the callbacks section of our configuration guide.
Fetch intron and upstream/downstream sequences
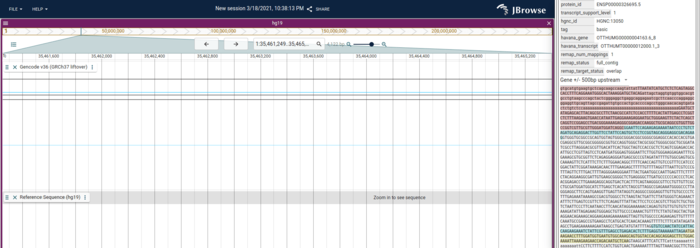
We also have several other improvements including the ability to get intron and upstream/downstream sequence in the feature details
Interactive documentation using Storybook
Another new update is the first release of our interactive Storybook docs for the embeddable React Linear Genome View. The docs contain live examples of how the LGV component can be used, along with source-code examples. The site can be found here.
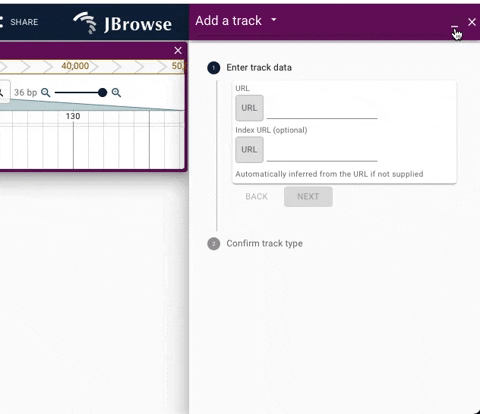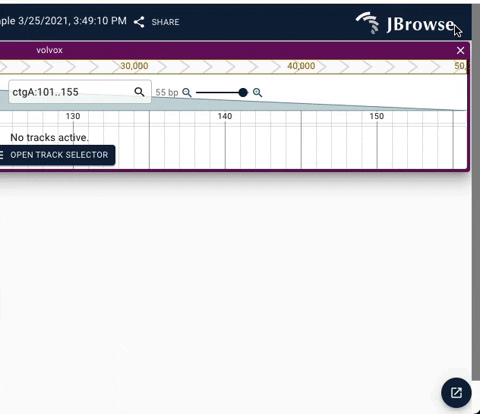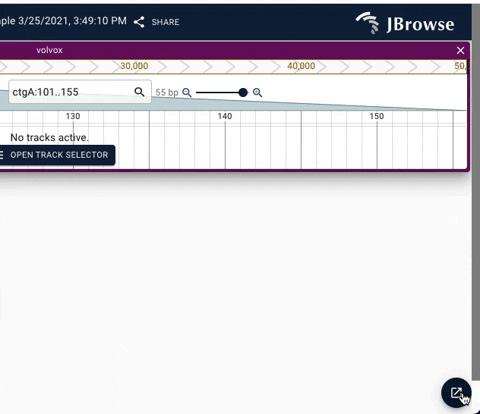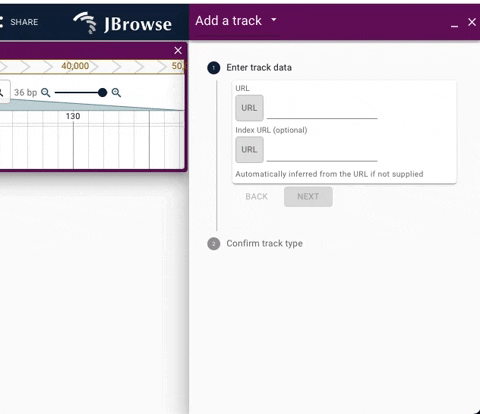
We have added a dropdown to enhance navigation between stack of active widgets. The update also adds a minimize button to allow quick access to full screen JBrowse web.
See below for demos of the new navigation UI.
Demo of using the minimize button in the drawer
Downloads
To install JBrowse 2 for the web, you can download the link above, or you can use the JBrowse CLI to automatically download the latest version. See the JBrowse web quick start for more details.
1.1.0 (2021-03-29)
🚀 Enhancement
core
- 1846 Improve copy+paste in the data grids for feature details (@cmdcolin)
- 1814 Add ability to get promoter sequence and intron sequence for genes from the feature details panel (@cmdcolin)
- 1816 Remove some animation effects (@cmdcolin)
- 1778 Adds dropdown to show drawer widget stack (@teresam856)
- 1685 Change callbacks language from JavaScript to Jexl (@peterkxie)
Other
- 1831 Add dialog for launching breakpoint split view from variant feature details (@cmdcolin)
- 1803 Transcript and gene glyphs can now display implied UTRs, active by default (@cmdcolin)
- 1808 Add another heuristic for returning gene features from BigBed (@cmdcolin)
- 1774 Add warning dialog in LGV before returning to import form to prevent accidentally losing the current view (@cmdcolin)
🐛 Bug Fix
core
- 1811 Check for existence of window more robustly to allow in SSR or node applications (@elliothershberg)
- 1793 Fix dotplot rendering outside it’s allowed bounds (@cmdcolin)
- 1783 Add hic aborting and fix remoteAbort signal propagation (@cmdcolin)
- 1723 A few bugfixes (@garrettjstevens)
Other
- 1815 Clear tracks when using “Return to import form” (@cmdcolin)
- 1819 Standardized sentence casing on drawer widget titles (@cmdcolin)
- 1796 Bump generic-filehandle for fixing CORS errors from Chrome cache pollution (@cmdcolin)
📝 Documentation
- 1824 Add storybook docs page for nextjs usage (@elliothershberg)
- 1770 1469 storybook deploy (@elliothershberg)
- 1807 Update developer guide to cover displays, and highlight working external plugins (@cmdcolin)
- 1779 Collaborative release announcement editing (@rbuels)
- 1791 Add a couple more demos for our live version with MDX (@cmdcolin)
🏠 Internal
Other
- 1820 Create v1.1.0.md, draft of release announcements (@garrettjstevens)
- 1823 Add note about previewing changelog to CONTRIBUTING.md (@garrettjstevens)
core
- 1834 Change jbrowse-components monorepo default branch from ‘master’ to ‘main’ (@rbuels)
Committers: 6
- Colin Diesh (@cmdcolin)
- Elliot Hershberg (@elliothershberg)
- Garrett Stevens (@garrettjstevens)
- Peter Xie (@peterkxie)
- Robert Buels (@rbuels)
- Teresa Martinez (@teresam856)
Posted to the GMOD News on 2021/03/30