Difference between revisions of "GBrowse"
(→On-line documentation) |
m (format changes) |
||
| Line 5: | Line 5: | ||
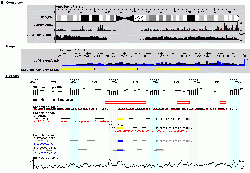
[[image:gbrowse_screenshot1.gif|right|thumb|250px|GBrowse running on [http://hapmap.org/downloads/index.html HapMap.org] [[Media:gbrowse_screenshot1.gif|View at full resolution]]]] | [[image:gbrowse_screenshot1.gif|right|thumb|250px|GBrowse running on [http://hapmap.org/downloads/index.html HapMap.org] [[Media:gbrowse_screenshot1.gif|View at full resolution]]]] | ||
| − | The Generic Genome Browser is a combination of database and interactive Web page for manipulating and displaying annotations on genomes. Some of its features: | + | The Generic Genome Browser (GBrowse) is a combination of database and interactive Web page for manipulating and displaying annotations on genomes. Some of its features: |
* Simultaneous bird's eye and detailed views of the genome. | * Simultaneous bird's eye and detailed views of the genome. | ||
| Line 13: | Line 13: | ||
* Order and appearance of tracks are customizable by administrator and end-user. | * Order and appearance of tracks are customizable by administrator and end-user. | ||
* Search by annotation ID, name, or comment. | * Search by annotation ID, name, or comment. | ||
| − | * Supports third party annotation using [ | + | * Supports third party annotation using [[GFF]] formats. |
* Settings persist across sessions. | * Settings persist across sessions. | ||
| − | * DNA and GFF dumps. | + | * DNA and [[GFF]] dumps. |
| − | * Connectivity to different databases, including [ | + | * Connectivity to different databases, including [[BioSQL]] and [[Chado]]. |
* Multi-language support. | * Multi-language support. | ||
* Third-party feature loading. | * Third-party feature loading. | ||
| Line 29: | Line 29: | ||
==Installation== | ==Installation== | ||
| − | GBrowse is Perl-based. It can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed. See the step-by-step instructions below for detailed instructions: | + | GBrowse is [[Glossary#Perl|Perl]]-based. It can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed. See the step-by-step instructions below for detailed instructions: |
* [[GBrowse_MacOSX_HOWTO|Install on MacOSX]] | * [[GBrowse_MacOSX_HOWTO|Install on MacOSX]] | ||
| Line 42: | Line 42: | ||
* [http://gmod.cvs.sourceforge.net/*checkout*/gmod/Generic-Genome-Browser/docs/tutorial/tutorial.html?pathrev=stable GBrowse Administration Tutorial] | * [http://gmod.cvs.sourceforge.net/*checkout*/gmod/Generic-Genome-Browser/docs/tutorial/tutorial.html?pathrev=stable GBrowse Administration Tutorial] | ||
* [[GBrowse_Install_HOWTO|GBrowse source code installation HOWTO]] | * [[GBrowse_Install_HOWTO|GBrowse source code installation HOWTO]] | ||
| − | * [[ | + | * [[GBrowse Configuration HOWTO]] |
* {{CVS|Generic-Genome-Browser/htdocs/general_help.html}} | * {{CVS|Generic-Genome-Browser/htdocs/general_help.html}} | ||
* {{CVS|Generic-Genome-Browser/htdocs/annotation_help.html}} | * {{CVS|Generic-Genome-Browser/htdocs/annotation_help.html}} | ||
| Line 54: | Line 54: | ||
* {{CVS|Generic-Genome-Browser/docs/pod/GENBANK_HOWTO.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/GENBANK_HOWTO.pod}} | ||
* {{CVS|Generic-Genome-Browser/docs/pod/PLUGINS_HOWTO.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/PLUGINS_HOWTO.pod}} | ||
| − | |||
* {{CVS|Generic-Genome-Browser/docs/pod/INSTALL.MacOSX.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/INSTALL.MacOSX.pod}} | ||
* {{CVS|Generic-Genome-Browser/docs/pod/DAS_HOWTO.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/DAS_HOWTO.pod}} | ||
| Line 61: | Line 60: | ||
* {{CVS|Generic-Genome-Browser/docs/pod/FAQ.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/FAQ.pod}} | ||
* {{CVS|Generic-Genome-Browser/docs/pod/MAKE_IMAGES_HOWTO.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/MAKE_IMAGES_HOWTO.pod}} | ||
| − | * {{CVS|Generic-Genome-Browser/docs/pod/README-gff-files.pod}} | + | * {{CVS|Generic-Genome-Browser/docs/pod/README-gff-files.pod}} (see also [[GFF]]) |
* {{CVS|Generic-Genome-Browser/docs/pod/GBROWSE_IMG.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/GBROWSE_IMG.pod}} | ||
* {{CVS|Generic-Genome-Browser/docs/pod/ORACLE_AND_POSTGRESQL.pod}} | * {{CVS|Generic-Genome-Browser/docs/pod/ORACLE_AND_POSTGRESQL.pod}} | ||
Revision as of 21:59, 30 December 2008
GBrowse[1] is the most popular viewer in GMOD. For a list of GBrowse and GMOD installations see the GMOD Users page. For a demo of its features, try the WormBase, FlyBase, or Human Genome Segmental Duplication Database web sites.
Contents
Description
The Generic Genome Browser (GBrowse) is a combination of database and interactive Web page for manipulating and displaying annotations on genomes. Some of its features:
- Simultaneous bird's eye and detailed views of the genome.
- Scroll, zoom, center.
- Use a variety of premade glyphs (see the BioPerl Glyph documentation for a list) or create your own.
- Attach arbitrary URLs to any annotation.
- Order and appearance of tracks are customizable by administrator and end-user.
- Search by annotation ID, name, or comment.
- Supports third party annotation using GFF formats.
- Settings persist across sessions.
- DNA and GFF dumps.
- Connectivity to different databases, including BioSQL and Chado.
- Multi-language support.
- Third-party feature loading.
- Customizable plug-in architecture (e.g. run BLAST, dump & import many formats, find oligonucleotides, design primers, create restriction maps, edit features)
GBrowse Versions
GBrowse 1.X (currently 1.69) is the stable series that has been in use since 2002. It is recommended for production use.
GBrowse 2.0 is a rewrite of the original GBrowse to add dynamic updating via AJAX and a smoother user experience. In addition, it provides administrators with the ability to attach a different genome database to each GBrowse track, making it much easier to manage and update tracks. It also provides a distributed backend system of "slave" renderers, allowing each track to be rendered in parallel on a different machine and significantly increasing performance. GBrowse 2.0 is currently in alpha test, but is quite usable. If you wish to try it, consult the GBrowse 2.0 HOWTO.
Installation
GBrowse is Perl-based. It can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed. See the step-by-step instructions below for detailed instructions:
- Install on MacOSX
- Install on Windows
- Install on Ubuntu and other Debian-based systems
- Install on Fedora Core and other RPM-based systems
- Install on Gentoo Linux system
- Source Code Install (for other Linux systems)
Documentation
On-line documentation
- GBrowse Administration Tutorial
- GBrowse source code installation HOWTO
- GBrowse Configuration HOWTO
- Generic-Genome-Browser/htdocs/general_help.html
- Generic-Genome-Browser/htdocs/annotation_help.html
- GBrowse Popup Balloons
- GBrowse FAQ
POD documentation
There are many useful POD documents included with the distribution. These are converted to HTML files when you install the package, and can be found in /gbrowse/docs/pod:
- Generic-Genome-Browser/docs/pod/BIOSQL_ADAPTER_HOWTO.pod
- Generic-Genome-Browser/docs/pod/GENBANK_HOWTO.pod
- Generic-Genome-Browser/docs/pod/PLUGINS_HOWTO.pod
- Generic-Genome-Browser/docs/pod/INSTALL.MacOSX.pod
- Generic-Genome-Browser/docs/pod/DAS_HOWTO.pod
- Generic-Genome-Browser/docs/pod/INSTALL.pod
- Generic-Genome-Browser/docs/pod/README-chado.pod
- Generic-Genome-Browser/docs/pod/FAQ.pod
- Generic-Genome-Browser/docs/pod/MAKE_IMAGES_HOWTO.pod
- Generic-Genome-Browser/docs/pod/README-gff-files.pod (see also GFF)
- Generic-Genome-Browser/docs/pod/GBROWSE_IMG.pod
- Generic-Genome-Browser/docs/pod/ORACLE_AND_POSTGRESQL.pod
- Generic-Genome-Browser/docs/pod/README-lucegene.pod
Since these are in Perl POD format these files may contain formatting code when viewed in a Web browser.
Downloads
Source Code Download (tar.gz file)
Download the source from the SourceForge download page.
Net-based Installer Script
The net installer script, located at the GBrowse CVS repository will automatically download and install GBrowse and its Perl libraries for you. See Installation for details on using this script.
CVS
There are many new features in the current development version which have not been released yet. To get the latest version, please use anonymous CVS. The recommended branch to use is stable which is more stable than HEAD. Here is the recipe:
% cvs -d :pserver:anonymous@gmod.cvs.sourceforge.net:/cvsroot/gmod login CVS password: <hit return> % cvs -d :pserver:anonymous@gmod.cvs.sourceforge.net:/cvsroot/gmod co -r stable Generic-Genome-Browser
Once you have successfully checked out the Generic-Genome-Browser distribution, you can simply perform a cvs update inside the directory to get recent changes.
You can also browse the GBrowse CVS.
About Databases
GBrowse has a flexible adaptor (yes, it is spelled that way and is not "adapter") system for running off various types of databases/sources. A common question is "which adaptor should I be using?" This attempts to answer that question.
| Adaptor | Other required software | Roughly how many users | Pros | Cons |
|---|---|---|---|---|
| Bio::DB::SeqFeature::Store (use bp_seqfeature_load.pl) | MySQL, PostgreSQL, SQLite, BerkeleyDB | Many and growing fast. | Roughly 4X faster than Bio::DB::GFF for the same data; designed to work with GFF3 | Developed for use with GFF3; about 2X slower than Bio::DB::GFF to load a database |
| Bio::DB::GFF (use bp_load_gff.pl, bp_bulk_load_gff.pl, bp_fast_load_gff.pl) | A relational database server: MySQL, PostgreSQL, Oracle, or BerkeleyDB | Lots! (Especially MySQL) | Quite fast; large user base; Have to use this if your data is in the (now deprecated) GFF2 format. | Does not work well with GFF3 formatted data |
| Bio::DB::Sam (available from CPAN) | SAMtools | Growing (particularly with GBrowse2) | Very fast access to NextGen sequencing data | Difficult to use with GBrowse 1.70 |
| Bio::DB::BigWig and Bio::DB::BigWigSet (available from CPAN) | UCSC Formats | Growing (particularly with GBrowse2) | Very fast access to data in bigWig format | Difficult to use with GBrowse 1.70 |
| Bio::DB::BigBed (available from CPAN) | UCSC Formats | Growing (particularly with GBrowse2) | Very fast access to data in bigBed format | Difficult to use with GBrowse 1.70 |
| Bio::DB::Das::Chado (available from CPAN) | PostgreSQL and a Chado schema | Relatively few due to the specialized nature of Chado | Allows 'live' viewing of the features in a Chado database | Slow compared to Bio::DB::GFF |
| Bio::DB::Das::BioSQL (available from CPAN) | MySQL and a BioSQL schema | Relatively few due to the small number of BioSQL users | Allows 'live' viewing of the features in a BioSQL database | Slow compared to Bio::DB::GFF |
| Memory (ie, flat file database using either Bio::DB::GFF or SeqFeature::Store) | None | For real servers, none | Easy for rapid development and testing | Very slow for more than a few thousand features |
| LuceGene | Lucene (searches indexed flat files) | Relatively few |
Email Threads
There have been some useful email threads on adaptor choices and tradeoffs.
- Memory Database, 2010/06
Contacts
Please report bugs to the SourceForge Bug Tracker.
Please send questions to the GBrowse mailing list.
References
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:12368253