NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
GBrowse syn AGS Tutorial
| {{#icon: GBrowse_syn_logo.png|GBrowse_syn|200|GBrowse_syn}} |
GBrowse_syn Session Arthropod Genomics Symposium 2011 |
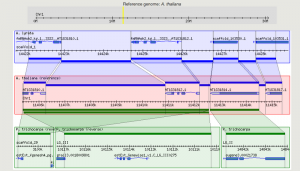
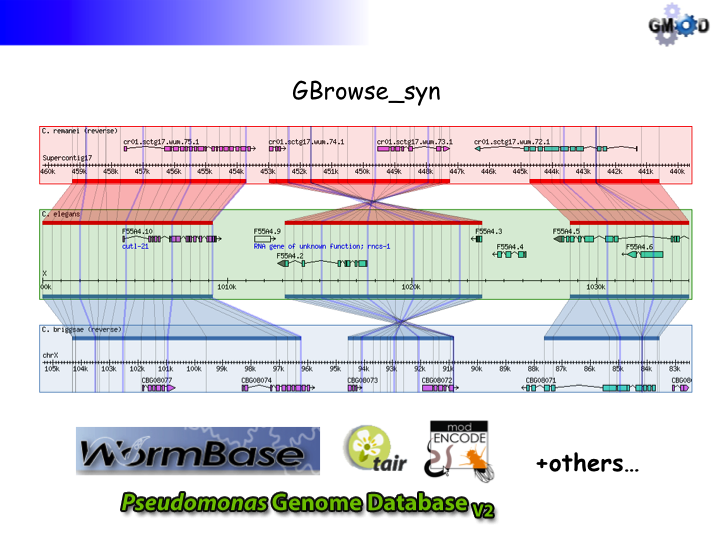
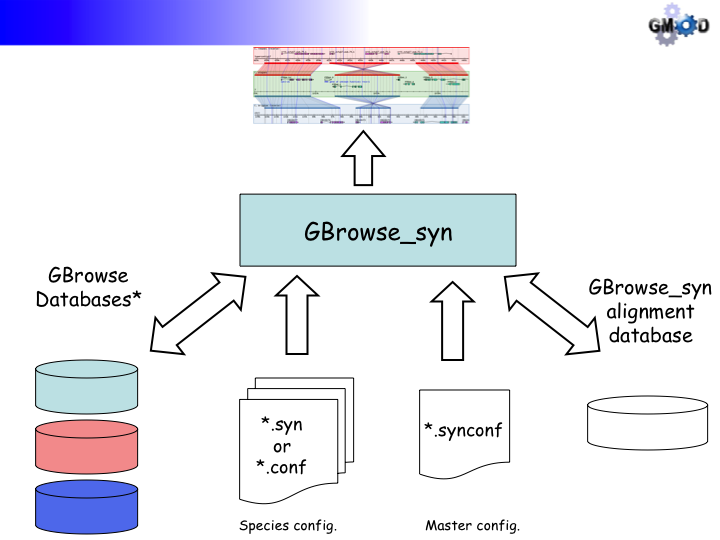
GBrowse_syn is a GBrowse-based synteny browser designed to display multiple genomes, with a central reference species compared to two or more additional species. It is included with the standard GBrowse package (version 1.69 and later).
Contents
GBrowse_syn Introduction
- An introductory talk will be presented using the slides below. Click the section to open.
Installing GBrowse_syn
GBrowse_syn is part of the GBrowse 2.0 package and will be pre-installed with GBrowse 2.0 installation.
$ sudo cpan -i Bio::Graphics::Browser2
Note: Gbrowse 2 has been pre-installed for this demonstration
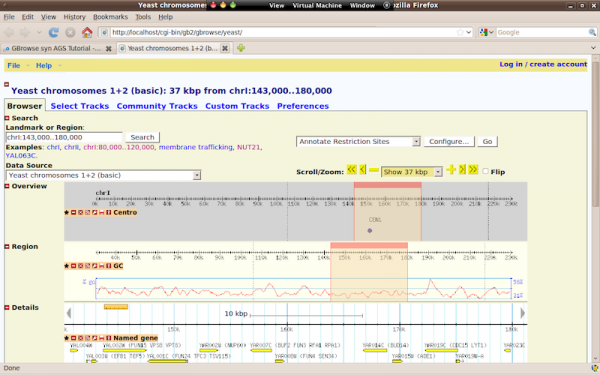
- Now point your browser to http://localhost/cgi-bin/gb2/gbrowse to see that GBrowse was installed correctly. You should see a page similar to the one below, which is the default yeast data source.
- Point your browser to http://localhost/cgi-bin/gb2/gbrowse_syn
Setting up the sample data
- Sample data and configuration information for GBrowse_syn come pre-packaged with GBrowse.
- The example we will use is a two-species comparison of rice (Oryza sativa) and one of its wild relatives*
*Data courtesy of Bonnie Hurwitz; sequences and names have been obfuscated to protect unpublished data
Setting up the Alignment Database
The alignment, or joining database will contain the sequence alignments between the two rice species. It will be in a MySQL database.
1) Create a MySQL database to hold the alignment data On the virtual disk, the MySQL root password is gmod
$ mysql -u root -p Enter password: **** Welcome to the MySQL monitor. Commands end with ; or \g. Your MySQL connection id is 37 Server version: 5.1.37-1ubuntu5.1 (Ubuntu)
Type 'help;' or '\h' for help. Type '\c' to clear the current input statement.
mysql> create database rice_synteny; Query OK, 1 row affected (0.00 sec)
mysql>
2) Give read-only (SELECT privileges in SQL) to the default apache user www-data. We can do this for all of the MySQL databases, since they are all for web applications
mysql> GRANT SELECT on *.* TO 'www-data'@'localhost'; Query OK, 0 rows affected (0.00 sec)
mysql> quit
3) Decompress the sample alignment data and load the database. You need to have root-level access (be a sudoer) for some of the steps below.
$ cd /var/lib/gbrowse2/databases/gbrowse_syn/alignments/ $ sudo gunzip rice.aln.gz
Have a look at the first few lines of the data:
$ head -20 rice.aln CLUSTAL W(1.81) multiple sequence alignment W(1.81)
rice-3(+)/16598648-16600199 ggaggccggccgtctgccatgcgtgagccagacggggcgggccggagacaggccacgtgg wild_rice-3(+)/14467855-14469373 gggggccgg------------------------------------agacaggccacgtgg ** ****** ***************
rice-3(+)/16598648-16600199 ccctgccccgggctgttgacccactggcacccctgtcccgggttgtcgccctcctttccc wild_rice-3(+)/14467855-14469373 ccctgccccgggctgttgacccactggcacccctgtcccgggttgtcgccctcctttccc ************************************************************
rice-3(+)/16598648-16600199 cgccatgctctaagtttgctcctcttctcgaacttctctctttgattcttcacgtcctct wild_rice-3(+)/14467855-14469373 cgccatgctctaagtttgctcctcttctcgaacttctctctttgattcttcacgtcctct ************************************************************
rice-3(+)/16598648-16600199 tggagcctccccttctagctcgatcacgctctgctcttccgcttggaggctggcaaaact wild_rice-3(+)/14467855-14469373 tggagcctccccttctagctcgatcgcgctctgctcttccgcttggaggctggcaaaact
The format is CLUSTALW. This is a formatting convention; it does not mean CLUSTALW was used to generate the alignment data. See Further Reading below for more information on data loading and the meta-data in the sequence names
4) Load the database with the script gbrowse_syn_load_alignments_msa.pl, which is automatically installed along with GBrowse. See the GBrowse_syn scripts page for details on the options for the script.
$ gbrowse_syn_load_alignments_msa.pl -u root -p gmod -d rice_synteny -c -v rice.aln
There are 1800 alignment blocks, so this will take a little while to run.
Setting up the Configuration Files
- The configuration files required for this data source are pre-installed with GBrowse, in /etc/gbrowse2/synteny/.
- There are two species' config files, rice_synteny.conf and wild_rice_synteny.conf, and the joining config file, oryza.synconf. The latter file has been disabled by appending a '.disabled' extension to the file name.
The joining config file, oryza.synconf:
[GENERAL]
description = BLASTZ alignments for Oryza sativa
===Sample Configuration Files===
# The synteny database
join = dbi:mysql:database=rice_synteny;host=localhost
# This option maps the relationship between the species data sources, names and descriptions
# The value for "name" (the first column) is the symbolic name that gbrowse_syn users to identify each species.
# This value is also used in two other places in the gbrowse_syn configuration:
# the species name in the "examples" directive and the species name in the .aln file
# The value for "conf. file" is the basename of the corresponding gbrowse .conf files.
# This value is also used to identify the species configuration stanzas at the bottom of the configuration file.
# name conf. file Description
source_map = rice rice_synteny "Domesic Rice (O. sativa)"
wild_rice wild_rice_synteny "Wild Rice"
tmpimages = /tmp/gbrowse2
imagewidth = 800
stylesheet = /gbrowse2/css/gbrowse_transparent.css
cache time = 1
config_extension = conf
# example searches to display
examples = rice 3:16050173..16064974
wild_rice 3:1..400000
zoom levels = 5000 10000 25000 50000 100000 200000 400000
# species-specific databases
[rice_synteny]
tracks = EG
color = blue
[wild_rice_synteny]
tracks = EG
color = red
A sample species config file, rice_synteny.conf:
[GENERAL]
description = Domestic rice chromosome 3
db_adaptor = Bio::DB::SeqFeature::Store
db_args = -adaptor memory
-dir /var/www/gbrowse2/databases/gbrowse_syn/rice
# Web site configuration info
tmpimages = /tmp/gbrowse2
[EG]
feature = gene:ensembl
glyph = gene
height = 10
bgcolor = peachpuff
fgcolor = hotpink
description = 0
label = 0
category = Transcripts
key = ensembl gene
balloon hover = Hello, my name is $name!
Note: the species databases are actually using the GFF3 flat file, in-memory adapter
Activating the Oryza Data Source
1) Make sure the temporary image directory specified in the config files exists and is world-writable
$ sudo mkdir /var/www/tmp $ sudo mkdir /var/www/tmp/gbrowse2 $ sudo chmod 777 /var/www/tmp/gbrowse2
2) Renaming the configuration file
$ cd /etc/gbrowse2/synteny $ sudo mv oryza.synconf.disabled oryza.synconf
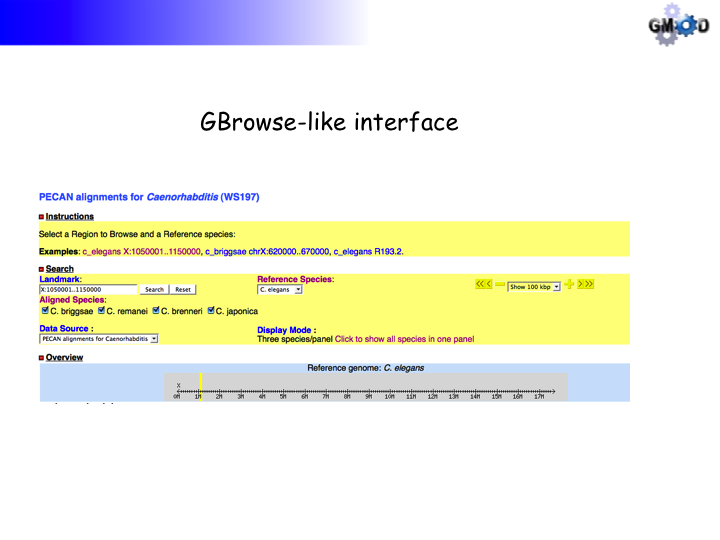
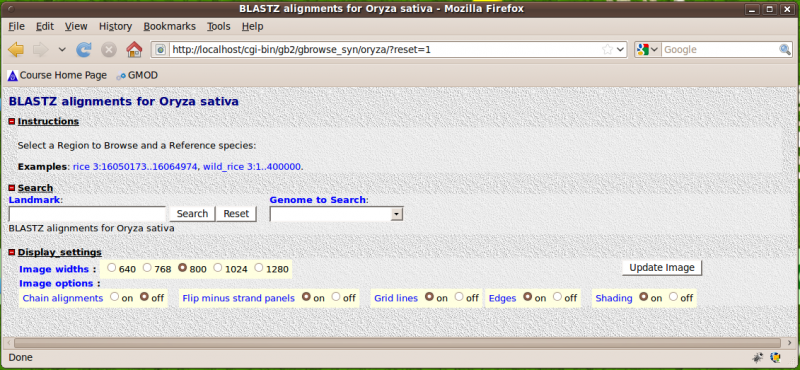
3) Point your browser to http://localhost/cgi-bin/gb2/gbrowse_syn/oryza. You should see:
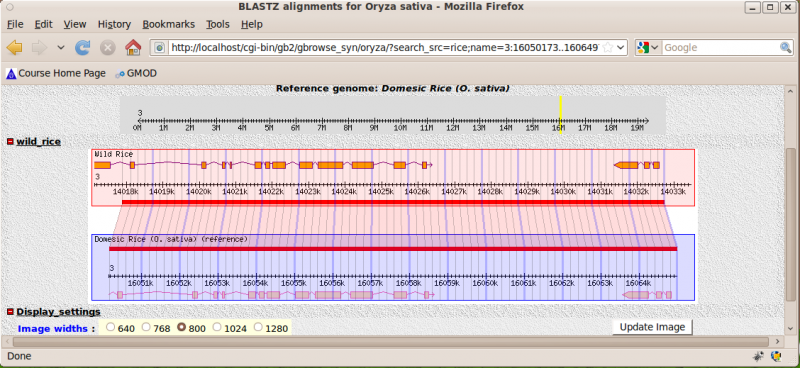
4) Click on the first example, you should (eventually) see:
Speeding up the Browser
You can speed up the image loading time by putting your species' GFF3 data into relational MySQL databases.
1) Create a database for each of the GFF data files (rice.gff3 and wild_rice.gff3).
$ mysql -uroot -pgmod
mysql> create database rice; Query OK, 1 row affected (0.00 sec)
mysql> create database wild_rice; Query OK, 1 row affected (0.00 sec)
mysql> quit
2) Populate the databases using the loading script bp_seqfeature_load.pl (pre-installed as part of BioPerl with GBrowse). This will load the GFF3 data into a MySQL relational database. Note the MySQL user will root-level privileges.
$ cd /var/www/gbrowse2/databases/gbrowse_syn/rice $ bp_seqfeature_load.pl -u root -p gmod -d rice -c -f rice.gff3 loading rice.gff3... Building object tree... 0.55s0s
Loading bulk data into database... 0.73s load time: 11.99s
$ cd ../wild_rice $ bp_seqfeature_load.pl -u root -p gmod -d wild_rice -c -f wild_rice.gff3 loading wild_rice.gff3... Building object tree... 0.55s7a
Loading bulk data into database... 0.69s load time: 12.02s
3) Modify the following stanza in the file rice_synteny.conf. This will convert your data source from a flat file database to a MySQL relational database.
# from
db_args = -adaptor memory
-dir /var/www/html/gbrowse/databases/gbrowse_syn/rice
# to
db_args = -dsn dbi:mysql:rice
4) repeat for wild_rice_synteny.conf
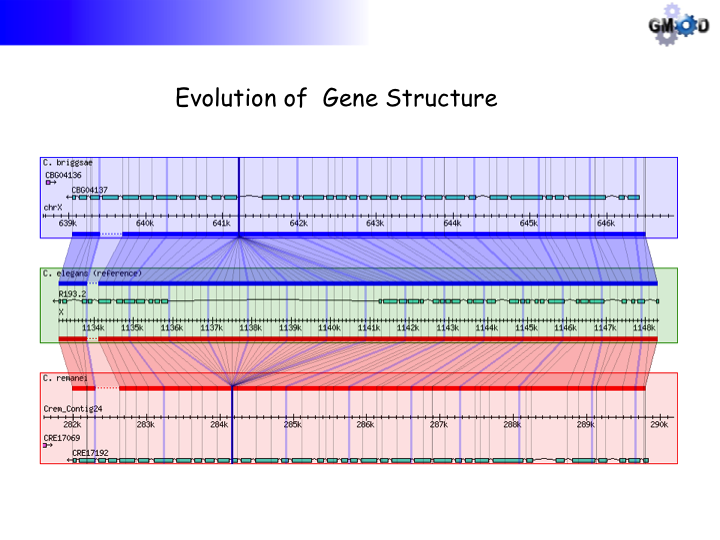
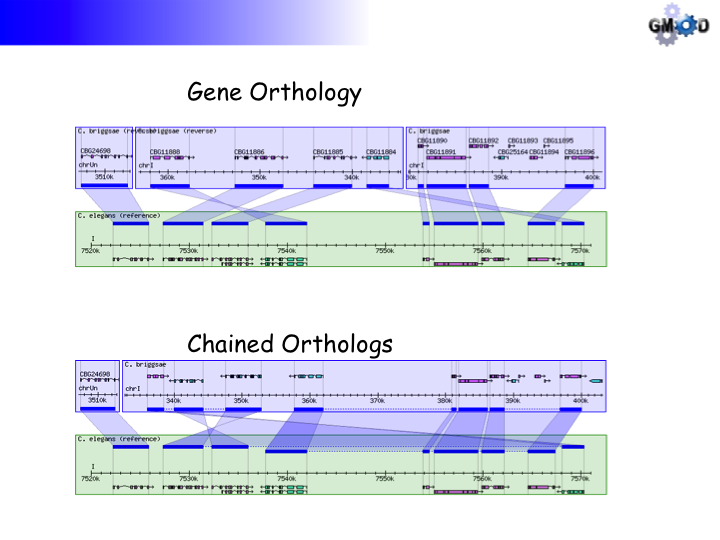
Using Non-alignment Data
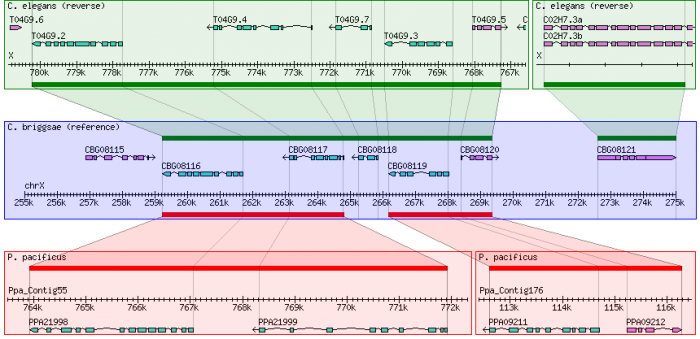
This example uses gene orthology-based synteny blocks* based created by OrthoCluster for three nematode species, C. elegans, C. briggsae and P. pacificus.
*Data courtesy of Jack Chen and Ismael Vergera
1) Download and unpack the data archive file orthocluster.tar.gz.
$ cd ~/Documents/Data/gbrowse_syn $ tar zxf orthocluster.tar.gz
2) Examine the contents of the ORTHOCLUSTER directory tree using the Unix tree command. It is not installed by default, so we will have to get it first.
$ sudo apt-get install tree [sudo] password for gmod: Reading package lists... Done Building dependency tree Reading state information... Done The following NEW packages will be installed: tree 0 upgraded, 1 newly installed, 0 to remove and 37 not upgraded. Need to get 31.1kB of archives. After this operation, 98.3kB of additional disk space will be used. Get:1 http://us.archive.ubuntu.com karmic/universe tree 1.5.2.2-1 [31.1kB] Fetched 31.1kB in 0s (37.0kB/s) Selecting previously deselected package tree. (Reading database ... 135915 files and directories currently installed.) Unpacking tree (from .../tree_1.5.2.2-1_i386.deb) ... Processing triggers for man-db ... Setting up tree (1.5.2.2-1) ...
Now we can use it
$ tree ORTHOCLUSTER/
ORTHOCLUSTER/
|-- conf
| |-- bri.conf
| |-- ele.conf
| |-- orthocluster.synconf
| `-- ppa.conf
|-- gff
| |-- bri.gff
| |-- ele.gff
| `-- ppa.gff
`-- orthocluster.txt
2 directories, 8 files
In the conf directory, there are configuration files for the joining database and each of the three species. They are similar in structure to the examples shown above, except that the database adapter Bio::DB::GFF and a gene aggregator are used because the GFF is version 2. For example:
[GENERAL]
description = C. briggsae
db_adaptor = Bio::DB::GFF
db_args = -dsn dbi:mysql:bri
# This is the GFF2 aggregator that assembles gene models
# from coding exon features with the same parent
aggregators = gene{coding_exon}
The gff directory contains gene annotations for each of the three species, derived from WormBase (release WS204). The files are in GFF2 format, which is why the Bio::DB::GFF adapter is required. A sample is shown here:
##gff-version 2 ##sequence-region I 1 15072421 ##sequence-region II 1 15279324 ##sequence-region III 1 13783685 ##sequence-region IV 1 17493784 ##sequence-region V 1 20924143 ##sequence-region X 1 17718854 I curated coding_exon 11641 11689 . + 0 CDS "Y74C9A.2" I curated coding_exon 14951 15160 . + 2 CDS "Y74C9A.2" I curated coding_exon 16473 16585 . + 2 CDS "Y74C9A.2" I curated coding_exon 43733 43961 . + 0 CDS "Y74C9A.1" I curated coding_exon 44030 44234 . + 2 CDS "Y74C9A.1" I curated coding_exon 44281 44324 . + 1 CDS "Y74C9A.1" I curated coding_exon 44372 44468 . + 2 CDS "Y74C9A.1" I curated coding_exon 44521 44677 . + 1 CDS "Y74C9A.1" I curated coding_exon 47472 47610 . + 0 CDS "Y48G1C.12" I curated coding_exon 47696 47858 . + 2 CDS "Y48G1C.12" I curated coding_exon 48348 48530 . + 1 CDS "Y48G1C.12" I curated coding_exon 49251 49416 . + 1 CDS "Y48G1C.12"
The file orthocluster.txt contains the synteny data. The first few lines are shown below. The first 12 fields in each row specify information about the synteny block in each species and the series of numbers are orthologous gene coordinate pairs that are used for linking orthologs with grid-lines in the GBrowse_syn display. See 'Alignment Data' under Further Reading below for more details of this loading format.
bri chrI 176154 183558 + . ppa Ppa_Contig88 27212 30786 + . 176154 27212 177594 30786 182118 27212 183558 30786 | 30786 183558 27212 182118 30786 177594 27212 176154 bri chrI 778780 799223 + . ppa Ppa_Contig88 533454 542961 - . 778780 539924 786778 542961 789497 533454 799223 538726 | 538726 799223 533454 789497 542961 786778 539924 778780 bri chrI 986150 994698 + . ppa Ppa_Contig77 29481 45600 - . 986150 37055 989649 45600 991428 29481 994698 36608 | 36608 994698 29481 991428 45600 989649 37055 986150 bri chrI 1453793 1461931 + . ppa Ppa_Contig132 156183 165414 - . 1453793 163110 1456404 165414 1456712 160849 1457637 162712 1458361 160204 1459245 160815 1459468 159346 1459854 160000 1459962 156183 1461931 159022 | 159022 1461931 156183 1459962 160000 1459854 159346 1459468 160815 1459245 160204 1458361 162712 1457637 160849 1456712 165414 1456404 163110 1453793
3) Set the $TMP environmental variable so that the database loading script knows where to put its temp files.
$ export TMP=/tmp
4) Create and load a Bio::DB:GFF database for C. elegans (ele). Use screen so that we can get the time-consuming loading script started and then use Ctrl-A D to set the screen running in the background and move on to other steps.
$ cd ORTHOCLUSTER/gff $ mysql -uroot -pgmod -e 'create database ele' $ screen bp_fast_load_gff.pl -u root -p gmod -d ele -c ele.gff
5) Repeat step 4 for the other two species (bri and ppa).
6) Create and load the alignment the alignment database. The gbrowse_syn_load_alignment_database.pl script is pre-installed with GBrowse.
$ cd .. $ mysql -uroot -pgmod -e 'create database orthocluster' $ gbrowse_syn_load_alignment_database.pl -u root -p gmod -d orthocluster -c -v orthocluster.txt
7) Copy the configuration files to the required location
$ cd conf $ sudo cp *conf /etc/gbrowse2/synteny
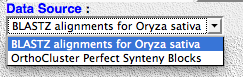
8) Go back to your browser and reload the rice page. There should now be a second data source in a pull-down menu.
9) Select the other data source and start browsing!
Mercator
MERCATOR facilitates whole-genome alignments using protein-coding exons as anchors in the alignment procedure. It creates a map of the synteny blocks among the genomes compared and can be used for pairwise or multi-way alignments. The nucleotide sequence for the synteny blocks is then aligned using alignment software (in this cased Mavid) This exampled assumes the MERCATOR pipeline has already been run to produce nucleotide alignment blocks. The results of MERCATOR include several files and directories. The necessary folder for this procedure is the alignments directory.
The typical directory structure includes an input and output directory. The output directory contains alignments; this contains all the data necessary for transformation to GBrowse syn. Example data are taken from pairwise alignments for the species Drosophila yakuba and D. melanogaster from the Web site http://www.biostat.wisc.edu/∼cdewey/fly CAF1. These data are the result of a MERCATOR and MAVID alignment between these two fly species. Although MAVID is used for the example data, other DNA sequence alignment software could be used on the synteny blocks identified by MERCATOR.
the map file contains locations of each synteny block in each of the genomes aligned in the order listed.
The file mercator.tab is in a tab delimited format designed for direct loading into the GBrowse syn alignment (or joining) database. The format has one tab-delimited record/line. Each line represents a synteny block, or alignment, with 13 fields: Reference Species Reference Seqid Reference Start Reference End Reference Strand Reference Cigar-string (not used; reserved for future use) Target Species Target Seqid Target Start Target End Target Strand Target Cigar-string (not used; reserved for future use)
Coordinate map (optional)
The coordinate map is used to save pair-wise nucleotide residue coordinates for
columns in the aligned sequences. It is not necessary to store coordinates for every
column. GBrowse syn usually uses multiples of 10, typically 100. The purpose of
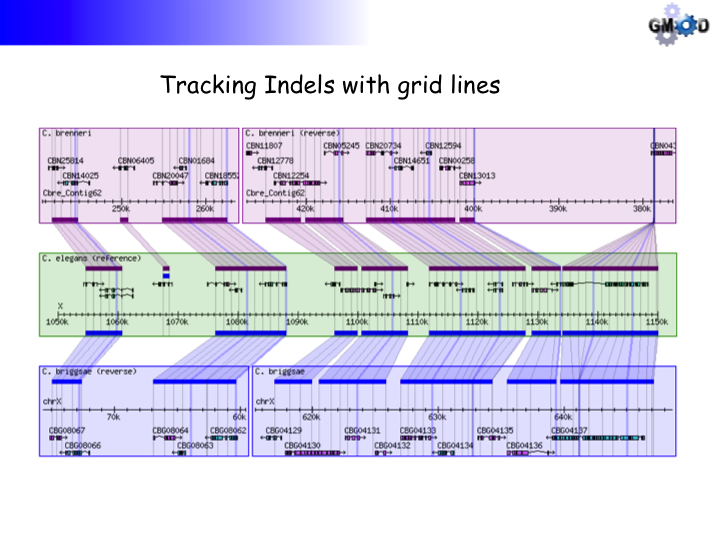
storing the coordinate information is to position grid lines in the graphical display
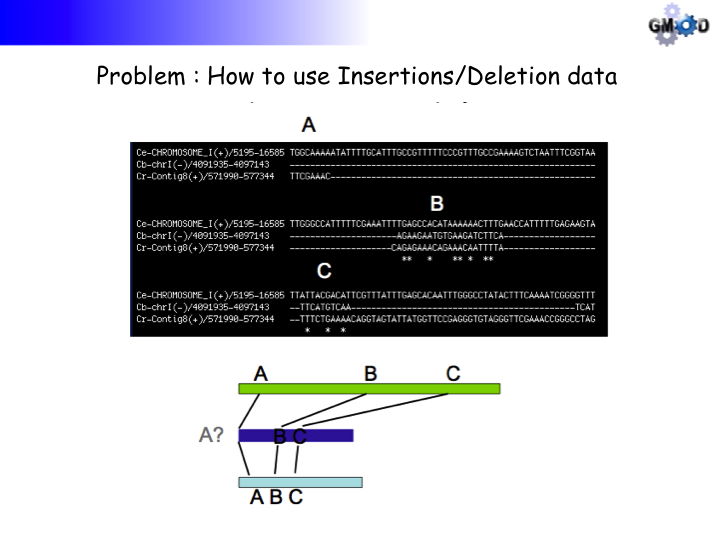
that will make large insertions and deletions in the sequences visible and intuitive.
The grid lines are equidistant on the reference sequence but can show insertions or
deletions by increasing or decreasing the distance between the lines, respectively, on
the target sequence. The format of field 13 (with spaces, not tabs):
rcoord1 tcoord1 rcoord2 tcoord2 | tcoorda rcoorda tcoordb rcoordb where: rcoordn reference nucleotide residue number n tcoordn target nucleotide residue number n n column in the alignment | Symbol delimiting reciprocal coordinate maps
NOTE: calculating the coordinate map is computationally intensive and the script may take a long time to run.
Load the GBrowse syn alignment database with the script gbrowse_syn_load alignment database.pl, which is preinstalled and can be run without specifying the path.
$ load alignment database.pl -u root -p gmod -d mercator -v -c mercator.txt
Further Reading
A Note on Whole Genome Alignments
The focus of the section of the course is on dealing with alignment or synteny data and using GBrowse_syn. However, how to generate whole genome alignments, identify orthologous regions, etc, are the subject of considerable interest, so some background reading is listed below:
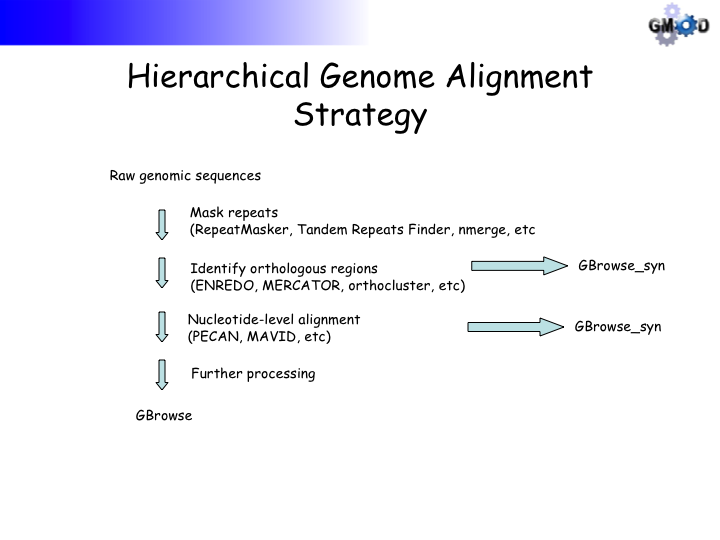
- Primer on Hierarchical Genome Alignment Strategies
- article on PECAN and ENREDO
- all about PECAN
- The gene annotations for each species are in GFF files.
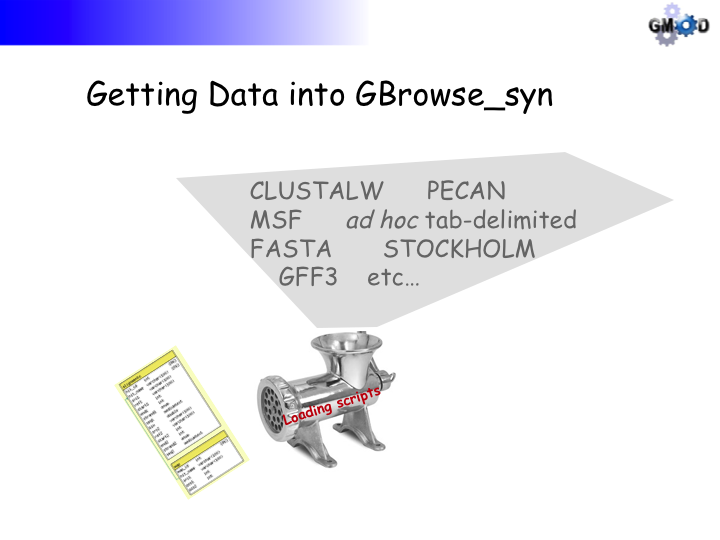
- The alignment data are in a constrained CLUSTALW format (They were not generated by the program CLUSTALW, which is not necessarily suitable for whole genome alignments)
- There are processing steps for the alignment data but it is very computationally intensive and we will load pre-processed data to get a head start.
Documentation
There is detailed documentation on the GMOD wiki for how to install, configure and use GBrowse_syn. To get started, browse these pages:
- GBrowse_syn overview
- Installation
- Configuration
- Alignment Data
- The user interface
- Presentations and workshops