NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Galaxy
- Mature release
- Development: active
- Support: active
Contents
About Galaxy
Galaxy is an open, web-based platform for accessible, reproducible, and transparent computational biomedical research.
- Accessibility: Galaxy enables users without programming experience to easily specify parameters and run tools and workflows.
- Reproducibility: Galaxy captures all information necessary so that any user can repeat and understand a complete computational analysis.
- Transparency: Galaxy enables users to share and publish analyses via the web and create Pages--interactive, web-based documents that describe a complete analysis.
Galaxy is open source for all organizations and is available on a wide variety of platforms and publicly accessible websites. Galaxy servers make analysis tools, genomic data, tutorial demonstrations, persistent workspaces, and publication services available to any scientist that has access to the Internet. Local Galaxy servers can be set up by downloading the Galaxy application and customizing it to meet particular needs.
2019 Galaxy Community Conference
The 2019 Galaxy Community Conference (GCC2019) will be held 1-6 July, in Freiburg Germany. Galaxy Community Conferences are an opportunity to participate in presentations, discussions, poster sessions, lightning talks and breakouts, all about high-throughput biology and the tools that support it. The 2019 conference includes training that offers in-depth topic coverage across several concurrent sessions, and a CollaborationFest.
Visit the Galaxy website.
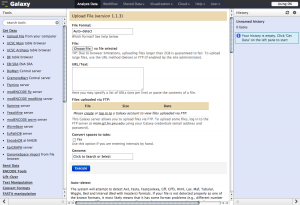
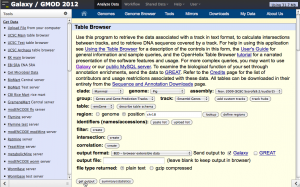
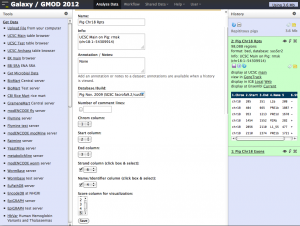
Screenshots
Downloads
Using Galaxy
Galaxy aims to be a zero configuration entirely self-contained system that provides a lightweight webserver, an embedded database and a multi-threaded job manager. All tools (and their parameters) can be specified via simple XML based configuration files.
Full documentation on all aspects of getting, installing, and using Galaxy is available from the Galaxy Community Hub.
Documentation
Publications, Tutorials, and Presentations
Publications on or mentioning Galaxy
See Citing Galaxy for a core list of 25+ papers on Galaxy, and the Galaxy publication library for a fuller list of papers mentioning or using Galaxy.
Tutorials
- A wealth of community created and maintained training materials are available:
Contacts and Mailing Lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| Galaxy (Search everything) |
galaxy-announce | Announcements of interest to the Galaxy community. Low volume and moderated. | Nabble, Mailman |
| Galaxy Help | General questions and discussion about using Galaxy. Also used for announcements relevant to the Galaxy user community. This is not a mailing list, but an online forum, based on the popular Discourse platform. High volume. | ||
| galaxy-dev | Discussion and questions regarding local installations and development of Galaxy. Medium volume. | Nabble, Mailman |
Galaxy in the wild
Public installations of Galaxy:
More on Galaxy
See Category:Galaxy
| Available on platform | web + |
| Has URL | http://getgalaxy.org/ +, https://github.com/galaxyproject/galaxy/ +, https://docs.galaxyproject.org +, https://galaxyproject.org/ +, https://twitter.com/galaxyproject +, https://www.zotero.org/groups/1732893/galaxy +, https://galaxyproject.org/use/ +, http://wiki.galaxyproject.org/PublicGalaxyServers + and https://galaxyproject.github.io + |
| Has description | Galaxy is an … Galaxy is an open, web-based platform for accessible, reproducible, and transparent computational biomedical research.
Galaxy is open source for all organizations and is available on a wide variety of platforms and publicly accessible websites. Galaxy servers make analysis tools, genomic data, tutorial demonstrations, persistent workspaces, and publication services available to any scientist that has access to the Internet. Local Galaxy servers can be set up by downloading the Galaxy application and customizing it to meet particular needs.
2019 Galaxy Community Conference[edit]The 2019 Galaxy Community Conference (GCC2019) will be held 1-6 July, in Freiburg Germany. Galaxy Community Conferences are an opportunity to participate in presentations, discussions, poster sessions, lightning talks and breakouts, all about high-throughput biology and the tools that support it. The 2019 conference includes training that offers in-depth topic coverage across several concurrent sessions, and a CollaborationFest.current sessions, and a CollaborationFest. + and Fully supported, publicly accessible platforms for using Galaxy + |
| Has development status | active + |
| Has input format | Roadmaps +, Sequences +, ab1 +, affybatch +, afg +, axt +, bam +, bcf +, bed +, bedgraph +, bigbed +, bigwig +, bowtie_base_index +, bowtie_color_index +, chrint +, cisml +, csfasta +, csv +, eland +, elandmulti +, encodepeak +, eset +, fasta +, fastq +, fastqcssanger +, fastqillumina +, fastqsanger +, fastqsolexa +, fli +, fped +, fphe +, fqtoc +, gatk_dbsnp +, gatk_interval +, gatk_recal +, gatk_report +, gatk_tranche +, gd_indivs +, gd_ped +, gd_sap +, gd_snp +, GFF +, GFF3 +, gtf +, interval +, lav +, ldindep +, len +, linecount +, lped +, maf +, malist +, memexml +, nex +, nhx +, pbed +, phyloxml +, picard_interval_list +, pileup +, pphe +, qual454 +, qualillumina +, qualsolexa +, qualsolid +, sam +, scf +, sff +, snpmatrix +, snptest +, tabular +, taxonomy +, twobit +, txt +, vcf +, velvet +, wig + and xml + |
| Has licence | Academic Free License 3.0 + |
| Has logo | GalaxyLogoBigger.png + |
| Has output format | zillions! + |
| Has software maturity status | mature + |
| Has support status | active + |
| Has title | Galaxy source code documentation +, Galaxy publication library +, ully supported, publicly accessible platforms for using Galaxy + and List of Galaxy Produced Software + |
| Has topic | Galaxy + |
| Is open source | Yes + |
| Link type | download +, source code +, documentation +, website +, social media +, other +, public server + and wild URL + |
| Release date | 2005 + |
| Tool functionality or classification | Genome Visualization and Editing +, Workflow Management +, Tool Integration + and Analysis + |
| Written in language | Python + and XML + |
| Has subobjectThis property is a special property in this wiki. | Galaxy#http://getgalaxy.org/ +, Galaxy#https://github.com/galaxyproject/galaxy/ +, Galaxy#https://docs.galaxyproject.org +, Galaxy#https://galaxyproject.org/ +, Galaxy#https://twitter.com/galaxyproject +, Galaxy#https://www.zotero.org/groups/1732893/galaxy +, Galaxy#https://galaxyproject.org/use/ +, Galaxy#http://wiki.galaxyproject.org/PublicGalaxyServers + and Galaxy#https://galaxyproject.github.io + |