NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Creating and Managing Subtracks with GBrowse2
For the main GBrowse 2.0 HOWTO article, see: GBrowse 2.0 HOWTO.
As of GBrowse version 2.09, you can create an unlimited number of subtracks within a single major track in order to group a series of datasets that are logically linked, such as a timecourse. You can choose which subtracks to show by default and the order in which they will appear. When the user clicks a designated area in the titlebar, a dialog box appears that allows the user to select which subtracks to make visible. The user can also drag subtrack labels up and down to adjust the order in which they are displayed.
There are two mechanisms for defining subtracks. The "Metadata" mechanism (new in version 2.48) is used when you have one feature per subtrack such as a whole genome quantitative ("wiggle") feature and each feature is enumerable by its display name. You provide a file that lists each feature subtrack explicitly.
The second mechanism is more flexible and is used when there are too many features to list explicitly and/or there are more than one feature per subtrack. In this scheme, each subtrack is defined by a set of feature filters. The filters are applied to each filter in turn, sorting them into the appropriate subtrack.
Contents
Using Metadata
If you have just a few features and there is a one-to-one correspondence between feature and subtrack, then the easiest way to define subtracks is by use of an external metadata file. A typical file looks like this:
[feature_name_1] :dbid = f101 :selected = 1 display_name = My First Feature type = some_type1 method = my_method1 source = my_source1 some_attribute = value1 another_attribute = value2 [feature_name_2] :dbid = f102 :selected = 1 display_name = My Second Feature type = some_type2 method = my_method2 source = my_source2 some_attribute = value3 another_attribute = value4 [feature_name_3] :dbid = f103 type = some_type2 method = my_method2 source = my_source2 some_attribute = value5 another_attribute = value6
Each [stanza] begins with the name of a feature as it is represented in the underlying database. Below each [stanza] heading are a series of tag=value pairs. The following tag names have special meaning:
:dbid Optional unique identifier for the subtrack; If provided, it can be used in the GBrowse
URL to select the subtrack.
:selected If true, this subtrack is selected by default when the containing track is turned on.
display_name Display name for the feature. If not present, will
default to the feature's native display name (i.e. the one in the [stanza]).
type What is returned by calling the feature's type() method.
method What is returned by calling the feature's method() method.
source What is returned by calling the feature's source() method.
score What is returned by calling the feature's score() method.
Any other tags become sortable attributes which are displayed by the GBrowse subtrack selection dialog box. For this to work properly, each tag must be present in each stanza. Tags that are present in some stanzas and not others are ignored.
Save this file anywhere convenient and then associate it with the desired track using the metadata option. This option takes the full path name to the metadata file. For example:
[ChIP-Seq] database = Peaks feature = signal glyph = vista_ploat metadata = /var/www/gbrowse2/databases/chip-seq/metadata.txt
Full Example
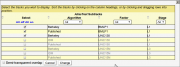
Here is a full working example of a metadata-based subtrack definition that shows a number of ChIP-seq experiments with the attributes "factor", "stage" and "algorithm". The subtrack selection dialog that this generates is shown on the right.
The database stanza:
[Chip:database] db_adaptor = Bio::DB::SeqFeature::Store db_args = -adaptor memory -dsn /var/www/gbrowse2/databases/elegans_peakcallcomparison search options = none
The features in this database are named "BLMP1-L1 Berkeley", "BLMP1-L1 IDR", etc. An excerpt from one of the GFF3 files that comprises this database can be found at Using the vista_plot Glyph.
The track stanza:
[ChIP-Seq] database = Chip feature = signal glyph = vista_ploat metadata = /var/www/gbrowse2/databases/chip-seq/metadata.txt
Metadata file in /var/www/gbrowse2/databases/chip-seq/metadata.txt.
[BLMP1-L1 Berkeley] :dbid=101 :selected=1 factor = BMLP1 stage = L1 algorithm = Berkeley [BLMP1-L1 IDR] :dbid=102 :selected=1 factor = BMLP1 stage = L1 algorithm = IDR [BLMP1-L1 Published] :dbid=103 factor = BMLP1 stage = L1 algorithm = Published [UNC130-L1 Berkeley] factor = UNC130 stage = L1 :dbid=104 algorithm = Berkeley [UNC130-L1 IDR] factor = UNC130 stage = L1 :dbid=105 algorithm = IDR [UNC130-L1 Published] factor = UNC130 stage = L1 :dbid=106 algorithm = Published
Save this file as /var/www/
Using Subtrack Select
This section describes how to use the subtrack select and subtrack table options to create subtracks based on filters.
A Basic Example
Here is a simple example to show how the system works. We start out with a gene track that has no subtracks:
[Genes] feature = gene glyph = gene database = sqlite-genes category = Genes:Coding label = 1 key = Wormbase Genes
The behavior of this track is to show both forward and reverse strand genes packed together for maximum efficiency, as shown in the figure on the right. Let's say we would prefer for forward and reverse stranded genes to be sorted into separate subtracks so that they do not intermingle. This can be done by adding subtrack select and subtrack table options to the configuration:
[Genes]
feature = gene
glyph = gene
database = sqlite-genes
category = Genes:Coding
label = 1
subtrack select = Strand strand
subtrack table = +1 ;
-1
key = Wormbase Genes
The "subtrack select" option defines a partitioning scheme for the data in the track. It consists of one or more lines defining the dimensions on which to partition the data. In this case, we are partitioning on only one dimension, the strandedness of the feature. The definition of this dimension is <Dimension Label> <method> (whitespace delimited, as usual), where the dimension label is a human-readable column label on the selection dialog and the method is any of the methods recognized by Bio::SeqFeatureI objects ("display_name", "primary_tag", "source_tag", "score", "has_tag", "tag_value", "start", "length", etc). In the example above, we are going to partition on the strand() method, and to label this dimension "Strand" on the popup menu presented to the user.
The "subtrack table" option defines the values on which to partition the data. In this case, we are going to partition the data into two subtracks: one for positive strand features (strand() returns +1) and one for negative strand features (strand() returns -1). This option has two or more lines, each one separated by a semicolon. Each line corresponds to a subtrack, which will be filtered by matching against the value(s) specified on the line. We simply list +1 and -1 as our two subtrack filter values.
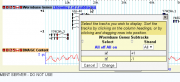
Reloading the browser now gives the track shown on the upper right. When the user clicks on the titlebar where it says "Showing 2 of 2 tracks", he can control the display of the subtracks using the dialog on the lower right.
This is good, but has two aesthetic issues. First, the labels on the subtracks appear as "+1" and "-1" which is not intuitive. Similarly, the strand values in the selection dialog also appear as +1 and -1. We can considerably improve this by attaching human-readable labels to the dimension values and subtracks. Here is an improved configuration file:
[Genes]
feature = gene
glyph = gene
database = sqlite-genes
category = Genes:Coding
label = 1
subtrack select = Strand strand
subtrack table = :Forward +1 ;
:Reverse -1
subtrack select labels = +1 "Forward Strand" ;
-1 "Reverse Strand"
key = Wormbase Genes
We've done two things here. First, we've modified the "subtrack table" option so that each line is preceded with :Name where Name is what we want to appear to the left of the subtrack in the display. The ":" symbol is required in front of the name but will not appear in the display. It can appear anywhere relative to the match items in the option.
Second, we added a "subtrack select labels" option to the stanza. This relabels the selectable dimension values with the desired human-readable labels within the dialog box itself. Notice that subtracks can have different names than their selection labels. In this example, we choose "Forward Strand" for +1 features in the dialog box, but "Forward" for the subtrack name.
The effect of these modifications are shown in the two figures to the right.
Multidimensional Subtracks
The previous example partitioned subtracks on a single dimension. This example will show how to create individually-selectable subtracks based on multiple selection dimensions. As an example, we use a track based on the modENCODE ChIP-seq tracks. These have four different dimensions, corresponding to the antibody used to bring down chromatin-bound transcription factors, the organism's developmental stage, the temperature at which the organism was grown, and whether the data set has been validated.
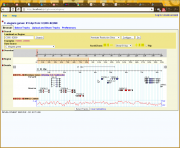
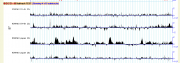
The images below and to the right show a development version of the histone modification ChIP-seq track from the modENCODE project and the dialog box used to select among the subtracks. This track was created with the following configuration:
[ChIP-seq]
subtrack select = Antibody tag_value antibody ;
Stage tag_value stage ;
Temperature tag_value temp ;
Confirmed has_tag confirmed ;
subtrack table = H3K4Me3 E0-4h 23 1 * ;
H3K4Me3 E4-8h 23 0 ;
H3K4Me3 pupae 23 0 ;
H3K4Me3 pupae 26 0 ;
H3K9Me2 E0-4h 23 0 * ;
H3K9Me2 E4-8h 23 1 ;
H3K9Me2 pupae 23 0 ;
H3K9Me2 pupae 26 1 ;
H3K27Me3 E0-4h 23 1 * ;
subtrack select labels = E0-4h "Early embryo" ;
E4-8h "Late embryo" ;
pupae "Pupating larvae" ;
brief comment = This track shows modENCODE ChIP-seq characterization of
histone marks across various stages and growth conditions.
In this case, the subtrack select option has four lines, each one separated by a semicolon (the semicolon on the final line is optional). Each line defines a subtrack dimension and has the format described above, consisting of a column label and a Bio::SeqFeature method call. However, in this case all the dimensions are contained in the features' tags (also known as feature "attributes"), which are accessed using the tag_value() and has_tag() methods. To specify which tags we are interested in, there is a third space-delimited argument that contains the tag name. So the Antibody dimension is determined by calling $feature->tag_value('antibody') and the Confirmed dimension is determined by calling $feature->has_tag('confirmed').
The subtrack table option lists each of the possible subtracks. The first four columns correspond to the four dimensions specified by subtrack select, e.g. antibody followed by developmental stage, followed by temperature and confirmation status. In this case we do not specify subtrack labels for reasons discussed later. However, since there are a lot of subtracks, we do not want them all to be displayed by default. The optional asterisk symbol (*) falling anywhere inside a subtrack table line indicates that this subtrack is to be turned on by default. If no asterisks are present, all subtracks are turned on.
The brief comment option serves to define a short description of the subtracks that will be printed in the subtrack selection dialog box.
Adding subtrack IDs to the Table
If you wish to embed GBrowse in another application, you will probably want to assign IDs to the subtracks so that they can be turned on and off via the GBrowse URL in the same way that whole tracks are turned on and off. To do this, append "=identifier" to each line of the subtrack table like this:
subtrack table = H3K4Me3 E0-4h 23 1 * =100 ;
H3K4Me3 E4-8h 23 0 =101 ;
H3K4Me3 pupae 23 0 =102 ;
H3K4Me3 pupae 26 0 =103 ;
H3K9Me2 E0-4h 23 0 * =104 ;
H3K9Me2 E4-8h 23 1 =105 ;
H3K9Me2 pupae 23 0 =106 ;
H3K9Me2 pupae 26 1 =107 ;
H3K27Me3 E0-4h 23 1 * =108 ;
The identifier can be any combination of letters and numbers. Its exact position on the line doesn't matter.
The identifiers can then be used to select subtrack in the GBrowse URL:
http://your.site/cgi-bin/gb2/gbrowse/elegans/?q=I:1000..2000;l=ChIP-seq/100+102+103
This will select the region between positions 1000 and 2000 on chromosome I, and turn on the ChIP-seq track, subtracks 100, 102 and 103.
Labeling Subtracks
Subtracks are labeled on the left side of the panel. If you are using "label_position=left" in your stanza, or the genes glyph with "label_transcripts" set to true, then it is possible for the labels of individual features to collide with the subtrack labels. There are two ways around this problem. One is to turn off subtrack labeling entirely and to let each individual feature's label identify subtracks. This works well with chromosome-wide features such as wiggle tracks when there is only one feature per subtrack (this is illustrated in the modENCODE data set above). The other is to relocate the position of the subtrack label to the top of the subtrack, where it won't clash with the feature labels. GBrowse will attempt to detect potential clash cases, and to configure them for you automatically, but there may be cases where you to have intervene manually.
To implement the first scheme, set the option "group_label" to a false value:
group_label = 0
Be sure that the label assigned to each feature is sufficiently informative that they can substitute for the subtrack label.
To implement the second scheme, set "group_label_position" to "top":
group_label_position = top
For compatibility with earlier behavior, GBrowse will automatically set group_label to 0 if the track consists of quantitative data (uses one of the wiggle, xyplot, density or whisker glyphs). Please set group_label to a true value in order to activate subtrack labeling.
Hiding Subtracks with no Data
By default, if a subtrack has no data in the currently displayed region, its label will still be printed to show that the subtrack exists. If you prefer, you can set hide empty subtracks to a true value, in which case the display of empty subtracks will be suppressed.