Difference between revisions of "GBrowse syn Database"
From GMOD
(→Database Loading Scripts) |
(→Database Loading Scripts) |
||
| Line 35: | Line 35: | ||
==Database Loading Scripts== | ==Database Loading Scripts== | ||
1) split large clustal files into one alignment/file with the script <span class="pops">[http://gmod.cvs.sourceforge.net/viewvc/*checkout*/gmod/Generic-Genome-Browser/bin/gbrowse_syn/split_clustal.pl?pathrev=stable split_clustal.pl]</span> | 1) split large clustal files into one alignment/file with the script <span class="pops">[http://gmod.cvs.sourceforge.net/viewvc/*checkout*/gmod/Generic-Genome-Browser/bin/gbrowse_syn/split_clustal.pl?pathrev=stable split_clustal.pl]</span> | ||
| + | |||
| + | gunzip my_huge_clustal_file.aln.gz | perl split_clustal.pl /path/to/smaller_alignment_files | ||
| + | |||
| + | 2) parse the alignments using the script [http://gmod.cvs.sourceforge.net/viewvc/*checkout*/gmod/Generic-Genome-Browser/bin/gbrowse_syn/clustal2hit.pl?pathrev=stable clustal2hits.pl] | ||
Revision as of 12:04, 14 January 2009
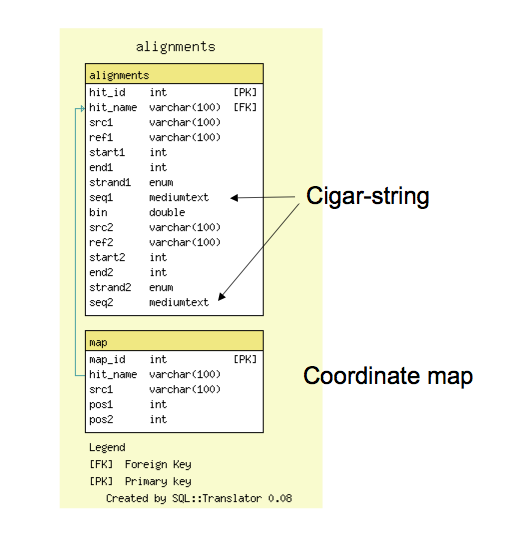
Schema
- The alignment database is very simple; it has a tables for all reciprocal 'hits', or alignment features and a table for (optional) 1:1 coordinate maps
- The alignments table contains coordinate information and also support cigar-line representations and the alignment to facilitate future reconstruction of the alignment within GBrowse_syn.
Loading
- The starting point for loading the alignment database is a CLUSTALW format multiple sequence alignment
- The source of the alignment data is up to you but the supported format for entry is currently CLUSTALW
Clustal alignment format
CLUSTAL W(1.81) multiple sequence alignment
c_briggsae-chrII(+)/43862-46313 ATGAGCTTCCACAAAAGCATGAGCTTTCTCAGCTTCTGCCACATCAGCATTCAAATGATC
c_remanei-Crem_Contig172(-)/123228-124941 ATGAGCCTCTACAACCGCATGATTCTTTTCAGCCTCTGCCACGTCCGCATTCAAATGCTC
c_brenneri-Cbre_Contig60(+)/627772-630087 ATGAGCCTCCACAACAGCATGATTTTTCTCGGCTTCCGCCACATCCGCATTCAAATGATC
c_elegans-II(+)/9706834-9708803 ATGAGCCTCTACTACAGCATGATTCTTCTCAGCTTCTGCAACGTCAGCATTCAGATGATC
****** ** ** * ****** ** ** ** ** ** ** ** ******* *** **
c_briggsae-chrII(+)/43862-46313 CGCACAAATATGATGCACAAATCCACAACCTAAAGCATCTCCGATAACGTTGACCGAAGT
c_remanei-Crem_Contig172(-)/123228-124941 AGCACAAATGTAATGAACGAATCCGCATCCCAACGCATCGCCAATCACATTCACAGATGT
c_brenneri-Cbre_Contig60(+)/627772-630087 CGCACAAATGTAGTGGACAAATCCGCATCCCAAAGCGTCTCCGATAACATTTACCGAAGT
c_elegans-II(+)/9706834-9708803 TGCACAAATGTGATGAACGAATCCACATCCCAATGCATCACCGATCACATTGACAGATGT
******** * ** ** ***** ** ** ** ** ** ** ** ** ** ** ** **
c_briggsae-chrII(+)/43862-46313 CCGGAGTCGATCCCTGAAT-----------------------------------------
c_remanei-Crem_Contig172(-)/123228-124941 ACGAAGTCGGTCCCTATAAGGTATGATTTTATATGA----TGTACCATAAGGAAATAGTC
c_brenneri-Cbre_Contig60(+)/627772-630087 ACGAAGTCGATCCCTGAAA---------TCAGATGAGCGGTTGACCA---GAGAACAACC
c_elegans-II(+)/9706834-9708803 ACGAAGTCGGTCCCTGAAC--AATTATTT----TGA----TATA---GAAAGAAACGGTA
** ***** ***** *
NOTE: The sequence naming convention "species-seqid(strand)/start-end" shown in the above example is essential for the data to be loaded correctly
Database Loading Scripts
1) split large clustal files into one alignment/file with the script split_clustal.pl
gunzip my_huge_clustal_file.aln.gz | perl split_clustal.pl /path/to/smaller_alignment_files
2) parse the alignments using the script clustal2hits.pl