Difference between revisions of "GBrowse syn Database"
From GMOD
(→Database Loading Scripts) |
(→Load the database) |
||
| Line 65: | Line 65: | ||
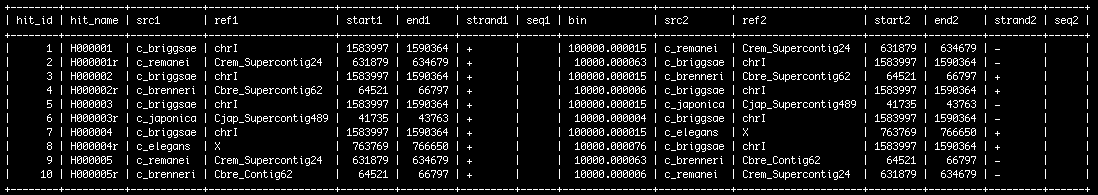
The alignment data | The alignment data | ||
[[Image:alignments_table.png]] | [[Image:alignments_table.png]] | ||
| − | + | ||
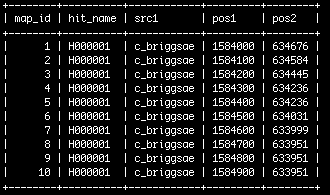
The grid mapping data | The grid mapping data | ||
[[Image:grid_map.png]] | [[Image:grid_map.png]] | ||
| + | |||
| + | <br clear="all"> | ||
Revision as of 12:36, 14 January 2009
Contents
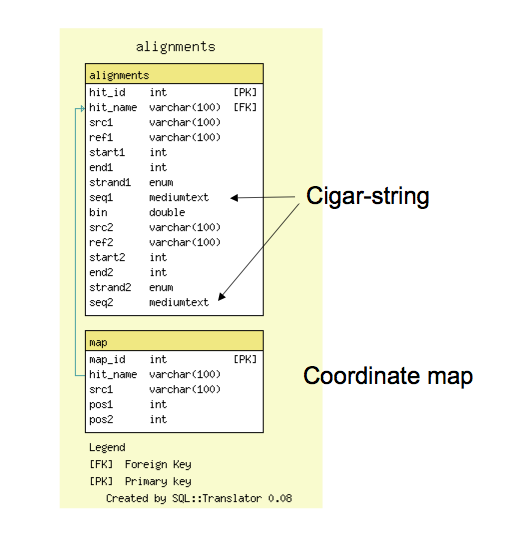
Schema
- The alignment database is very simple; it has a tables for all reciprocal 'hits', or alignment features and a table for (optional) 1:1 coordinate maps
- The alignments table contains coordinate information and also support cigar-line representations and the alignment to facilitate future reconstruction of the alignment within GBrowse_syn.
Loading
- The starting point for loading the alignment database is a CLUSTALW format multiple sequence alignment
- The source of the alignment data is up to you but the supported format for entry is currently CLUSTALW. This format is not mandatory but the parsing script below will need to be adjusted to support other formats
Clustal alignment format
CLUSTAL W(1.81) multiple sequence alignment
c_briggsae-chrII(+)/43862-46313 ATGAGCTTCCACAAAAGCATGAGCTTTCTCAGCTTCTGCCACATCAGCATTCAAATGATC
c_remanei-Crem_Contig172(-)/123228-124941 ATGAGCCTCTACAACCGCATGATTCTTTTCAGCCTCTGCCACGTCCGCATTCAAATGCTC
c_brenneri-Cbre_Contig60(+)/627772-630087 ATGAGCCTCCACAACAGCATGATTTTTCTCGGCTTCCGCCACATCCGCATTCAAATGATC
c_elegans-II(+)/9706834-9708803 ATGAGCCTCTACTACAGCATGATTCTTCTCAGCTTCTGCAACGTCAGCATTCAGATGATC
****** ** ** * ****** ** ** ** ** ** ** ** ******* *** **
c_briggsae-chrII(+)/43862-46313 CGCACAAATATGATGCACAAATCCACAACCTAAAGCATCTCCGATAACGTTGACCGAAGT
c_remanei-Crem_Contig172(-)/123228-124941 AGCACAAATGTAATGAACGAATCCGCATCCCAACGCATCGCCAATCACATTCACAGATGT
c_brenneri-Cbre_Contig60(+)/627772-630087 CGCACAAATGTAGTGGACAAATCCGCATCCCAAAGCGTCTCCGATAACATTTACCGAAGT
c_elegans-II(+)/9706834-9708803 TGCACAAATGTGATGAACGAATCCACATCCCAATGCATCACCGATCACATTGACAGATGT
******** * ** ** ***** ** ** ** ** ** ** ** ** ** ** ** **
c_briggsae-chrII(+)/43862-46313 CCGGAGTCGATCCCTGAAT-----------------------------------------
c_remanei-Crem_Contig172(-)/123228-124941 ACGAAGTCGGTCCCTATAAGGTATGATTTTATATGA----TGTACCATAAGGAAATAGTC
c_brenneri-Cbre_Contig60(+)/627772-630087 ACGAAGTCGATCCCTGAAA---------TCAGATGAGCGGTTGACCA---GAGAACAACC
c_elegans-II(+)/9706834-9708803 ACGAAGTCGGTCCCTGAAC--AATTATTT----TGA----TATA---GAAAGAAACGGTA
** ***** ***** *
NOTE: The sequence naming convention "species-seqid(strand)/start-end" shown in the above example is essential for the data to be loaded correctly
Database Loading Scripts
Split large clustal files into smaller ones
Split large clustal files into one alignment/file with the script split_clustal.pl. This is necessary for very large files that would otherwise overload the bioperl alignment parser.
gunzip my_huge_clustal_file.aln.gz | perl split_clustal.pl /path/to/smaller_alignment_files
Parse the alignments
Parse the alignments using the script clustal2hits.pl
perl clustal2hit.pl /path/to/smaller_alignment_files/*.aln |gzip -c >processed_alignments.txt.gz
- The output of this script is an ad hoc intermediate format that encodes the alignment coordinates plus 1:1 mapping of coordinates within the alignment (to facilitate accurate grid-lines in GBrowse_syn)
- Each record is a sinle line (wrapped here for display only)
#species1 seqid1 start1 end1 strand1 reserved species2 seqid2 start2 end2 strand2 reserved \ # pos1-1 pos1-2 ... posn-1 posn-2 | pos1-2 pos1-1 ... posn-2 posn-1 c_briggsae chrI 1583997 1590364 + . c_remanei Crem_Contig24 631879 634679 - . \ 1584000 634676 1584100 634584 (truncated for display...) | 631900 1590333 632000 1590233 (truncated for display...)
Create a mysql database
mysql -uroot -ppassword -e 'create database my_database'
Load the database
load the database with the script load_alignment_database.pl
gunzip -c processed_alignments.txt.gz | perl load_alignment_database.pl
- Once loaded, the data look something like this: