Difference between revisions of "GBrowse img"
DanielRenfro (Talk | contribs) |
DanielRenfro (Talk | contribs) m |
||
| Line 5: | Line 5: | ||
__TOC__ | __TOC__ | ||
==Description== | ==Description== | ||
| − | + | This CGI script is an interface to the Generic Genome Browser for the purpose of retrieving dynamic images of a region of the genome. It can be used as the destination of an <img> tag like this: | |
<img src="http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?name=NC_000913:3923000..3951000"> | <img src="http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?name=NC_000913:3923000..3951000"> | ||
| − | The script can also be used to superimpose one or more external features onto the display, for example for the purpose of displaying BLAST hits, an STS or a knockout in the context of the genome. | + | The script can also be used to superimpose one or more external features onto the display, for example for the purpose of displaying BLAST hits, an STS or a knockout in the context of the genome. It is designed to use a URL-based API to draw or embed [[GBrowse]] images without loading the full genome browser interface. gbrowse_img can be used to embed Gbrowse images in other webpages or to create high-resolution images appropriate for publications. |
==Examples== | ==Examples== | ||
| Line 33: | Line 33: | ||
DH10B | DH10B | ||
MG1655 | MG1655 | ||
| − | . | + | ''etc.'' |
| − | == | + | ===Listing Types=== |
| + | To get a list of available tracks (the '''type''' parameter), set the '''list''' parameter to ''types'' like so: | ||
| + | http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?list=types | ||
| + | {{Problem| | ||
| + | The format of this output is cryptic. Looks like: | ||
| + | # the track name | ||
| + | # text description | ||
| + | # whether this track is turned on by default or not | ||
| − | + | Someone who knows should probably edit this. | |
| − | + | --[[User:DanielRenfro|DanielRenfro]] 19:23, 27 April 2011 (UTC) | |
| + | }} | ||
| − | + | ## Feature types for source MG1655 | |
| − | + | Genes Genes default | |
| − | + | TranslationF 3-frame translation (forward) | |
| + | TranslationR 3-frame translation (reverse) | ||
| + | DNA DNA/GC Content | ||
| + | Protein | ||
| + | rRNA_Operons | ||
| + | RegulonDBtu RegulonDB txn units default | ||
| + | Cryptic_Prophage cryptic prophage default | ||
| + | ''etc.'' | ||
| − | |||
| − | |||
| − | == | + | ==CGI arguments== |
| − | + | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<a href="http://www.wormbase.org/db/seq/gbrowse_img/elegans?list=types">types</a> | <a href="http://www.wormbase.org/db/seq/gbrowse_img/elegans?list=types">types</a> | ||
| Line 93: | Line 88: | ||
==CGI arguments== | ==CGI arguments== | ||
| − | |||
The script recognizes the following CGI arguments, which can be passed either as GET or POST argument=value pairs. Argument pairs must be separated by semicolons (preferred) or by ampersands. Many of the options have one-letter aliases that can be used to reduce URL lengths. | The script recognizes the following CGI arguments, which can be passed either as GET or POST argument=value pairs. Argument pairs must be separated by semicolons (preferred) or by ampersands. Many of the options have one-letter aliases that can be used to reduce URL lengths. | ||
| Line 122: | Line 116: | ||
The arguments are explained in more detail here: | The arguments are explained in more detail here: | ||
| − | ; name | + | ; name / q |
: This argument specifies the region of the genome to be displayed. Several forms are recognized: | : This argument specifies the region of the genome to be displayed. Several forms are recognized: | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | :* ''name=Landmark'' | |
| − | + | :*: Display the landmark named "Landmark". Valid landmark names include chromosomes, contigs, clones, STSs, predicted genes, and any other landmark that the administrator has designated. Be careful when fetching large landmarks such as whole chromosomes! | |
| − | + | ||
| − | + | :* ''name=Landmark:start..end'' | |
| − | + | :*: Display the region between ''start'' and ''end'' relative to "Landmark". | |
| − | + | ||
| − | + | :* ''name=Class:Landmark'' | |
| − | + | :*: Display "Landmark", restricting to a particular class, such as "PCR_Product". The list of classes is under the control of the database administrator and is not yet available through this interface. | |
| − | + | ||
| − | + | :* ''name=Class:Landmark:start..end'' | |
| + | :*: As above, but restricted to the designated range. | ||
| + | |||
| + | If you use multiple <b>name</b> options, then this script will generate an overview image showing the position of each landmark. The alias "q" can be used to shorten the length of the URL. | ||
| Line 311: | Line 302: | ||
image instead. You may need to adjust the width and height attributes | image instead. You may need to adjust the width and height attributes | ||
in order to avoid browsers placing scrollbars around the frame. | in order to avoid browsers placing scrollbars around the frame. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==Known Bugs== | ||
| + | The cookie that stores the configuration options for plugins does not transfer from gbrowse to gbrowse_img, so tracks generated by annotation plugins, such as the Restriction site annotator, will not display correctly when the image URL is generated on one machine and then viewed on another. Uploaded files will transfer correctly, however. | ||
| + | |||
| + | ==Author== | ||
| + | [mailto: lstein@cshl.org Lincoln Stein] | ||
| + | Copyright (c) 2002-2004 Cold Spring Harbor Laboratory | ||
| + | |||
| + | This library is free software; you can redistribute it and/or modify | ||
| + | it under the same terms as Perl itself. | ||
| + | |||
| + | ==Mediawiki Extension== | ||
| + | The [http://www.mediawiki.org/wiki/GBrowseImage GbrowseImage extension] for Mediawiki will display an image (rendered by gbrowse_img) in a wiki-page. | ||
| + | |||
| + | ==See Also== | ||
| + | * [[Gbrowse]] | ||
[[Category:Documentation]] | [[Category:Documentation]] | ||
[[Category:GBrowse]] | [[Category:GBrowse]] | ||
[[Category:GMOD Components]] | [[Category:GMOD Components]] | ||
Revision as of 19:23, 27 April 2011
This page or section needs to be edited. Please help by editing this page to add your revisions or additions.
gbrowse_img - CGI script to generate genome images via the Generic Genome Browser
Contents
Description
This CGI script is an interface to the Generic Genome Browser for the purpose of retrieving dynamic images of a region of the genome. It can be used as the destination of an <img> tag like this:
<img src="http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?name=NC_000913:3923000..3951000">
The script can also be used to superimpose one or more external features onto the display, for example for the purpose of displaying BLAST hits, an STS or a knockout in the context of the genome. It is designed to use a URL-based API to draw or embed GBrowse images without loading the full genome browser interface. gbrowse_img can be used to embed Gbrowse images in other webpages or to create high-resolution images appropriate for publications.
Examples
The Escherichia coli K-12 MG1655 genome (from bp 3923000 to 3950999) has been used to generate the following figures. Also note that the URLs have been encoded.
Simple Example
http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?name=NC_000913:3923000..3951000;l=Genes%1Elandmarks:overview%1EGenes:region%1Elandmarks:region;width=400;format=GD;
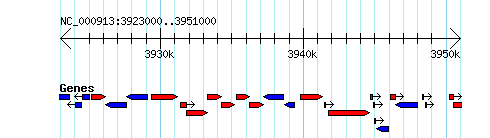
More complex
http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?name=NC_000913:3923000..3951000;l=Genes%1EDNA%1ERegulonDBtu%1ESigma70%1Elandmarks:overview%1EGenes:region%1Elandmarks:region;width=800;id=20736090abb824610d9d3bc89c8b4256;format=GD;keystyle=between;grid=1
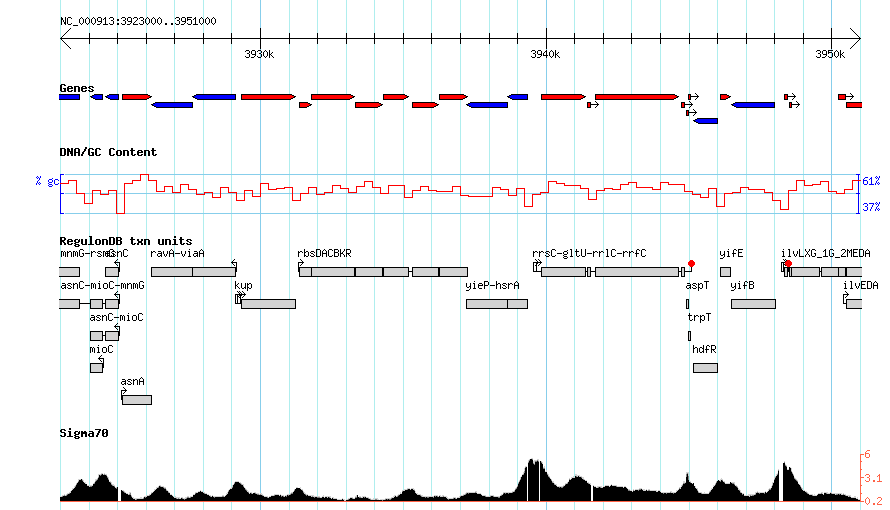
Listing Sources
You can get a list of sources (genomes, chromosomes, etc.), by setting the list parameter to sources like so:
http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?list=sources
## Sources ATCC_8739 BL21 BL21_DE3 BL21_Gold_DE3 BW2952 DH10B MG1655 etc.
Listing Types
To get a list of available tracks (the type parameter), set the list parameter to types like so:
http://heptamer.tamu.edu/cgi-bin/gb2/gbrowse_img/MG1655/?list=types
| Problem/Question: The format of this output is cryptic. Looks like:
Someone who knows should probably edit this. --DanielRenfro 19:23, 27 April 2011 (UTC) |
## Feature types for source MG1655 Genes Genes default TranslationF 3-frame translation (forward) TranslationR 3-frame translation (reverse) DNA DNA/GC Content Protein rRNA_Operons RegulonDBtu RegulonDB txn units default Cryptic_Prophage cryptic prophage default etc.
CGI arguments
<a href="http://www.wormbase.org/db/seq/gbrowse_img/elegans?list=types">types</a>
Will return this document: ## Feature types for source wormbase tRNA tRNAs NG Named Genes default CG Curated genes default PG Predicted genes WABA Briggsae alignments (WABA) ESTB ESTs aligned by BLAT (best) ESTO ESTs aligned by BLAT (other) mRNAB mRNAs aligned by BLAT (best) mRNAO mRNAs aligned by BLAT (other) RNAi RNAi experiments EXPR Expression chip profiles WTP Worm Transcriptome Project genes SNP SNPs TcI Transposon Insertions
CGI arguments
The script recognizes the following CGI arguments, which can be passed either as GET or POST argument=value pairs. Argument pairs must be separated by semicolons (preferred) or by ampersands. Many of the options have one-letter aliases that can be used to reduce URL lengths.
| Argument | Alias | Description |
|---|---|---|
| name | q | genomic landmark or range |
| type | t | tracks to include in image |
| width | w | desired width of image |
| options | o | list of track options (compact, labeled, etc) |
| abs | b | display position in absolute coordinates |
| add | a | added feature(s) to superimpose on the image |
| style | s | stylesheet for additional features |
| keystyle | k | where to place the image key |
| overview | force an overview-style display | |
| flip | f | flip image left to right |
| grid | turn grid on (1) or off (0) | |
| embed | generate full HTML for image and imagemap for use in an embedded frame | |
| format | format for the image (use "SVG" for scaleable vector graphics) | |
| list | get certain types of configuration information | |
| source | database name |
The arguments are explained in more detail here:
- name / q
- This argument specifies the region of the genome to be displayed. Several forms are recognized:
- name=Landmark
- Display the landmark named "Landmark". Valid landmark names include chromosomes, contigs, clones, STSs, predicted genes, and any other landmark that the administrator has designated. Be careful when fetching large landmarks such as whole chromosomes!
- name=Landmark
- name=Landmark:start..end
- Display the region between start and end relative to "Landmark".
- name=Landmark:start..end
- name=Class:Landmark
- Display "Landmark", restricting to a particular class, such as "PCR_Product". The list of classes is under the control of the database administrator and is not yet available through this interface.
- name=Class:Landmark
- name=Class:Landmark:start..end
- As above, but restricted to the designated range.
- name=Class:Landmark:start..end
If you use multiple name options, then this script will generate an overview image showing the position of each landmark. The alias "q" can be used to shorten the length of the URL.
- type (Alias
- t)
- This argument lists the feature types to display. The value of this argument is a list of track names separated by spaces ("+" characters when L-escaped). For example:
<img src="http://www.wormbase.org/db/seq/gbrowse_img/elegans?name=mec-3&type=tRNA+NG+WABA+CG+ESTB">
Multiple type= arguments will be combined to form a single space-delimited list.
The alias "t" can be used to shorten the length of the URL.
If the track name has a space in it, put quotes around the name:
type="microbe tRNA"+NG+WABA+CG+ESTB
<p>
options=tRNA+3+NG+3+WABA+1
<p>
The alias "o" can be used to shorten the length of the URL.
<p>
add=Landmark+Type+Name+start..end,start..end,start..end
"Landmark" is the landmark name, "Type" is a descriptive type that will be printed
in the image caption, "Name" is a name for the feature to be printed above it,
and start..end is a comma-delimited list of ranges for discontinuous feature.
Names that contain white space must be quoted, for example "BLAST hit".
Note that this all has to be URL-escaped, so an additional
feature named "Your Sequence", type "Blast Hit", that is located on chromosome III
in a gapped range between 20000 and 22000, will be formatted as:
<p>
add=III+%22Blast%20Hit%22+%22Your%20Sequence%22+20000..21000,21550..22000
<p>
One or both of the type and name can be omitted. If omitted, type will
default to "Your Features" and the name will default to "Feature XX" where
XX is an integer. This allows for a very simple feature line:
add=III+20000..21000,21550..22000
<p>
Multiple add= arguments are allowed. The alias "a" can be used to
shorten the length of the URL.
<p>
For example, if you have added a "Blast Hit" annotation, then you can tell the
renderer to use a red arrow for this glyph in this way:
style=%22Blast%20Hit%22+glyph=arrow+fgcolor=red
<p>
Mnemonic <tab> Full description of feature <tab> [default]
The third column contains the word "default" if the track will be shown by default when no type argument is provided. <p>
</pre>h_feat=SKT5@blue
</pre>
You may omit "@color", in which case the highlight will default to
yellow. You can specify multiple h_feat arguments in order to
highlight several features with distinct colors.
</pre>h_region=Chr3:200000..250000@wheat
</pre>
You may omit "@color", in which case the highlighted region
will default to
lightgrey. You can specify multiple h_region arguments in order to
highlight several regions with distinct colors.
</dl>
Putting it all together, here's a working (very long) URL:
<a href="http://www.wormbase.org/db/seq/gbrowse_img/elegans?name=B0001;add=B0001+pcr+pcr1+20000..333000;add=B0001+%22cool%20knockout%22+kn2+30000..20000,10000..5000;type=add+CG+WTP;style=pcr+glyph=primers;style=%22cool%20knockout%22+glyph=transcript2+bgcolor=orange;abs=1">http://www.wormbase.org/db/seq/gbrowse_img/elegans?name=B0001;add=B0001+pcr+pcr1+20000..333000;add=B0001+%22cool%20knockout%22+kn2+30000..20000,10000..5000;type=add+CG+WTP;style=pcr+glyph=primers;style=%22cool%20knockout%22+glyph=transcript2+bgcolor=orange;abs=1</a>
If you wish to associate the image with an imagemap so that clicking
on a feature takes the user to the destination configured in the
gbrowse config file, you may do so by placing the URL in an
<iframe> section and using the embed=1 flag:
<iframe src="http://localhost/cgi-bin/gbrowse_img/elegans?name=B0001;embed=1" width="100%" height="250"> <img src="http://localhost/cgi-bin/gbrowse_img/elegans?name=B0001"/> </iframe>
Placing an <img> tag inside the <iframe> tag arranges for older browsers that don't know about iframes to display the static image instead. You may need to adjust the width and height attributes in order to avoid browsers placing scrollbars around the frame.
Known Bugs
The cookie that stores the configuration options for plugins does not transfer from gbrowse to gbrowse_img, so tracks generated by annotation plugins, such as the Restriction site annotator, will not display correctly when the image URL is generated on one machine and then viewed on another. Uploaded files will transfer correctly, however.
Author
[mailto: lstein@cshl.org Lincoln Stein] Copyright (c) 2002-2004 Cold Spring Harbor Laboratory
This library is free software; you can redistribute it and/or modify it under the same terms as Perl itself.
Mediawiki Extension
The GbrowseImage extension for Mediawiki will display an image (rendered by gbrowse_img) in a wiki-page.