Difference between revisions of "CMap"
From GMOD
m |
m |
||
| Line 1: | Line 1: | ||
| − | {{ImageCenter| | + | {{ImageCenter|CMapLogo.png|CMap|180|}} |
__NOTITLE__ | __NOTITLE__ | ||
Revision as of 05:58, 8 July 2010
__NOTITLE__
Status
- Mature release
- Active development
- Active support
Resources
CMap is a web-based tool that allows users to view comparisons of genetic and physical maps. The package also includes tools for curating map data.
Contents
Requirements
- Perl 5.6.0 or higher
- A web server with CGI capabilites (e.g., Apache 1.x or 2.x)
- A relational database, e.g. MySQL (4.0+), Sybase Adaptive Server Enterprise, PostgreSQL, SQLite or Oracle (9.x)
- libgd 2.0.28 and GD.pm 2.15
- Various Perl modules available on CPAN
Documentation
- Gramene CMap User Tutorial,
- Available in both PDF and PowerPoint formats.
- CMap_API (SVN)
- README
- Changes
- INSTALL.pod
- ADMINISTRATION.pod
- CODE_OVERVIEW.pod
- TODO
- attributes-and-xrefs.pod
- CMAP Schema
- CMAP Tutorial at Texas A & M
- CMap Version 2 Design
Also see the CMap FAQ.
Presentations
- Genome Visualization and Comparison using CMap and Circos, poster from Genome Informatics 2009
- The CMap Comparative Map Browser and Displaying Population distributions with GBrowse and PhyloGeoViz from the GMOD Workshop at SMBE 2009
- CMap and CMAE from November 2007 GMOD Meeting
- CMap Version 0.16 from June 2006 GMOD Meeting
- CMap Update 2005 from May 2005 GMOD Meeting
- Gramene Comparative Genome Views: How Can Gramene Leverage Rice For The Other Grasses? poster form 2005 Plant and Animal Genomics meeting.
- CMap Update 2003 from September 2003 GMOD Meeting.
- CMap: Visualization of Comparative Maps from May 2003 GMOD Meeting.
Downloads
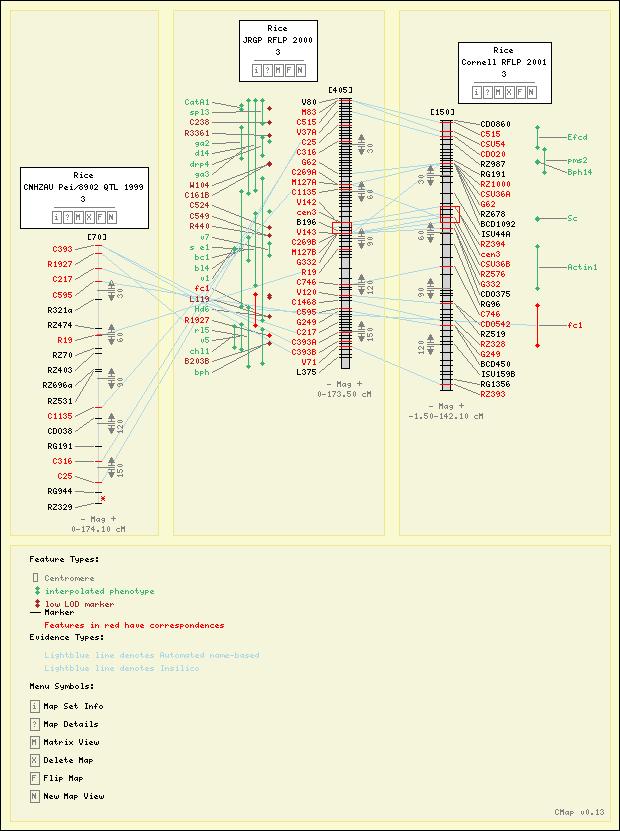
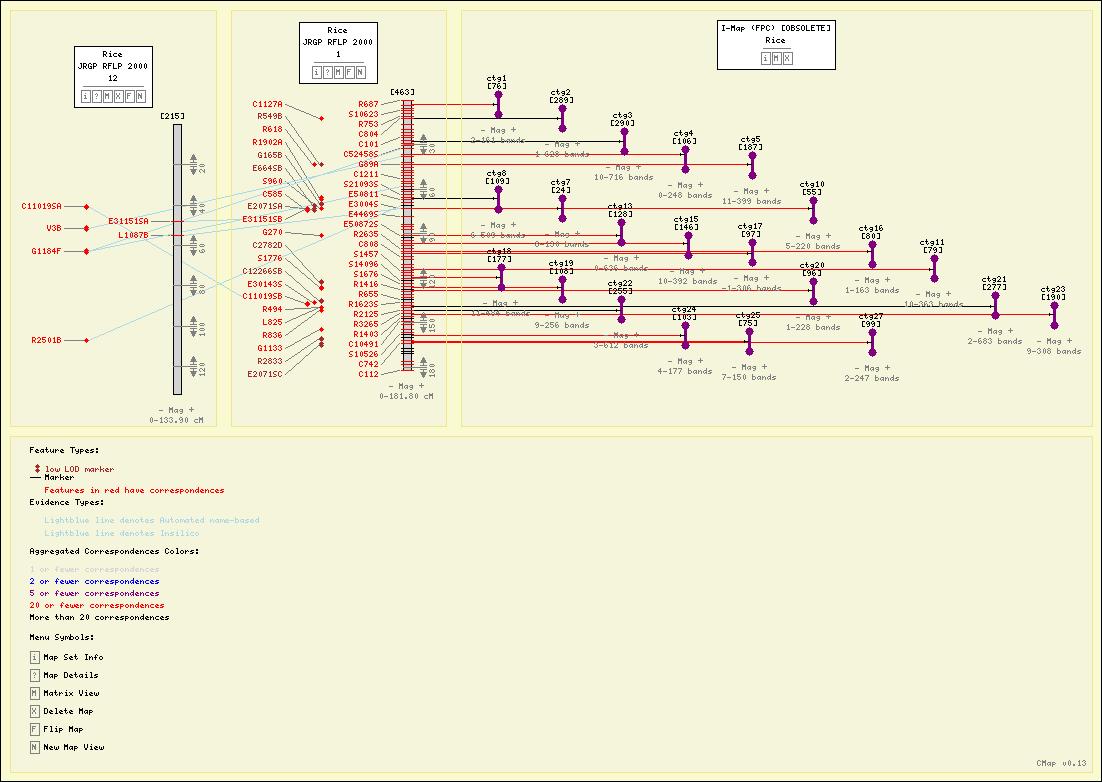
Demo & Screenshots
See the Gramene Project for a production version.
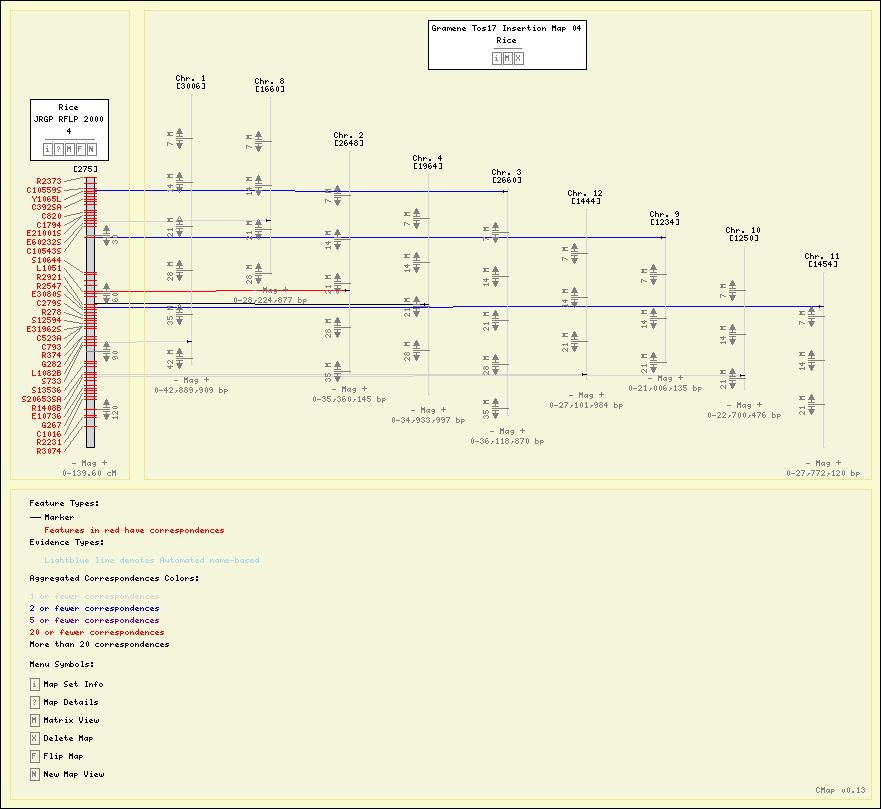
Screenshots
Here are some sample maps from the Gramene Project.
Sites Using CMap
The following is a partial list of groups using CMap.
- Gramene: Comparative mapping resource for grasses (plant)
- NCGR's Legume Information System: A publicly accessible legume resource that will integrate genetic and molecular data from multiple legume species and enable cross-species comparisons
- BeeBase: The Honey Bee genome database
- BASC: Brassica database
- GrainGenes
- MAGI
- Genome Database for Rosaceae
- Cotton Marker Database
- SCRI Plant Bioinformatics Group
- Pristionchus pacificus Physical Map
- Comparitive Location Database
- Soybean Breeders Toolbox
- Composite Wheat Map
- PeanutMap
- CottonDB
- TropGene
Code
Patches
- See the SourceForge Patch List.
- 1.0: Patched schema creation file for Postgres: {{SF_SVN|/cmap/trunk/sql/cmap.create.postgresql cmap.create.postgresql].
Mailing Lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| CMap | gmod-cmap | Discussion of CMap development, installation problems, etc. | Nabble (2010/05+), Sourceforge |
| gmod-cmap-commits | Notification of SVN activity for CMap. | Sourceforge |