JBrowse Configuration Guide
This page provides a comprehensive reference guide for installing and configuring JBrowse. If you find something that is missing or inaccurate, please feel very free to edit it!
Contents
- 1 Installation
- 2 Reference Sequences
- 3 Feature Tracks
- 4 Alignment Tracks (BAM)
- 5 Wiggle Tracks
- 6 Variant Tracks (VCF)
- 7 Feature Coverage Tracks
- 8 Image Tracks
- 9 Name Searching and Autocompletion
- 10 Removing Tracks
- 11 Compressing data on the server
- 12 Controlling JBrowse with the URL Query String
- 13 Data Export
- 14 Track Metadata
- 15 Faceted Track Selector
- 16 Dataset Selector
- 17 Anonymous Usage Statistics
- 18 Advanced Topics
- 18.1 Data Format Specification: LazyNCList Feature Store
- 18.2 Data Format Specification: Fixed-Resolution Tiled Image Store
- 18.3 Other Global Configuration
- 18.4 Using JBrowse with Existing Databases
- 18.5 Using JBrowse with Existing Web Services
- 18.6 Writing JBrowse-compatible Web Services
- 18.7 Writing JBrowse Plugins
- 19 External Links
Installation
At the most basic level, setting up JBrowse consists of:
- Placing a copy of the JBrowse directory somewhere in the web-servable part of your server's file system (often
/var/wwwby default) - Running the JBrowse setup script to install a few server-side dependencies
- Running one or more server-side scripts to create a directory containing a JBrowse-formatted copy of your data.
Both the JBrowse code and these data files must be in a location where the web server can serve them to users. Then, a user pointing their web browser at the appropriate URL for the index.html file in the JBrowse directory will see the JBrowse interface, including sequence and feature tracks reflecting the data source.
Reference sequence data should be added first (using prepare-refseqs.pl`), followed by annotation data. Once all of annotation data has been added, use generate-names.pl to make the feature names searchable.
Making a New JBrowse
0. Optionally, install libpng, Zlib, and GD development libraries, plus make and a C compiler. These are only needed for legacy wig-to-json.pl support. On Ubuntu, you could do this with:
sudo apt-get install libpng-dev libgd2-noxpm-dev build-essential
1. Download JBrowse onto your web server.
2. Unpack JBrowse into a directory that is served by your web browser. On many systems, this defaults to /var/www.
cd /var/www unzip JBrowse-*.zip
Make sure you have permissions to write to the contents of the jbrowse/ directory you have just created.
3. Run the automated-setup script, ./setup.sh, which will attempt to install all of JBrowse's (modest) prerequisites for you in the jbrowse/ directory itself. Note that setup.sh does not need to be run as root or with sudo.
4. Visit http://your.machine.address/jbrowse/index.html?data=sample_data/json/volvox. If you can see the included Volvox example data, you are ready to configure JBrowse to show your own data! The Getting Started with JBrowse Tutorial provides a very basic step-by-step guide to formatting your own data, and in-depth configuration reference information can be found on this page.
Upgrading an Existing JBrowse
To upgrade an existing JBrowse (1.3.0 or later) to the latest version, simply move its data directory (and jbrowse_conf.json if you are using it) into the directory of a newer JBrowse, and the new JBrowse will display that data.
To upgrade a 1.2.x JBrowse, copy its data directory into the new JBrowse directory, and point your browser at compat_121.html in the new JBrowse directory, instead of index.html. Or, if desired, you could simply overwrite index.html with compat_121.html.
If you are upgrading from a version of JBrowse older than 1.2.0, a fresh installation is required.
Reference Sequences
The reference sequences on which the browser will display annotations, and which provide a common coordinate system for all tracks. At a close enough zoom level, the sequence bases are visible in the "Reference Sequence" track.
The exact interpretation of "reference sequence" will depend on how you are using JBrowse. For a model organism genome database, each reference sequence would typically represent a chromosome or a contig. Before any feature or image tracks can be displayed in JBrowse, the reference sequences must be defined using the prepare-refseqs.pl formatting tool.
Reference Sequence Selector Configuration
JBrowse displays a dropdown selector for changing reference sequences.
For JBrowse versions 1.6.3 to 1.8.1, if more than 30 reference sequences are loaded, this selector is not shown by default. To force JBrowse to show the reference sequence dropdown selector, set refSeqDropdown: true in the configuration. This can be done in any configuration file, e.g. index.html, jbrowse_conf.json, or data/trackList.json.
In JBrowse version 1.9.0 and later, the reference sequence dropdown menu is always displayed. However, if there are too many reference sequences to practically display in the dropdown selector, the first portion of the sequence list is shown in the dropdown selector, along with a message stating that there are more reference sequences than can be shown. The maximum number of reference sequences in the selector is set by the refSeqSelectorMaxSize configuration variable, and defaults to 30.
In addition, beginning in JBrowse 1.9.0, the ordering of reference sequences in the selector is configurable using the refSeqOrder configuration variable. Supported values for this value are 'name', 'name descending', 'length', and 'length descending'. One instance in which this is particularly useful is
in displaying annotations on early-stage, incomplete genomic
assemblies: to display the N biggest contigs in the assembly in the
reference sequence selector dropdown, one can set refSeqOrder to
'length descending', and set refSeqSelectorMaxSize to N.
prepare-refseqs.pl
This script is used to format sequence data for use by JBrowse, and must be run before adding other tracks. In addition to formatting the sequence data, this script creates a track called "DNA" that displays the reference sequence. The simplest way to use it is with the --fasta option, which uses a single sequence or set of reference sequences from a FASTA file:
bin/prepare-refseqs.pl --fasta <fasta file> [options]
If the file has multiple sequences (e.g. multiple chromosomes), each sequence will become a reference sequence by default. You may switch between these sequences by selecting the sequence of interest via the pull-down menu to the right of the large "zoom in" button.
You may use any alphabet you wish for your sequences (i.e., you are not restricted to the nucleotides A, T, C, and G; any alphanumeric character, as well as several other characters, may be used). Hence, it is possible to browse RNA and protein in addition to DNA. However, some characters should be avoided, because they will cause the sequence to "split" - part of the sequence will be cut off and and continue on the next line. These characters are the hyphen and question mark. Unfortunately, this prevents the use of hyphens to represent gaps in a reference sequence.
In addition to reading from a fasta file, prepare-refseqs.pl can read sequences from a gff file or a database. In order to read fasta sequences from a database, a config file must be used.
Syntax used to import sequences from gff files:
bin/prepare-refseqs.pl --gff <gff file with sequence information> [options]
Syntax used to import sequences with a config file:
bin/prepare-refseqs.pl --conf <config file that references a database with sequence information> --[refs|refid] <reference sequences> [options]
| Option | Value |
|---|---|
| fasta, gff, or conf | Path to the file that JBrowse will use to import sequences. With the fasta and gff options, the sequence information is imported directly from the specified file. With the conf option, the specified config file includes the details necessary to access a database that contains the sequence information. Exactly one of these three options must be used. |
| out | A path to the output directory (default is 'data' in the current directory) |
| seqdir | The directory where the reference sequences are stored (default: <output directory>/seq) |
| noseq | Causes no reference sequence track to be created. This is useful for reducing disk usage. |
| refs | A comma-delimited list of the names of sequences to be imported as reference sequences. This option (or refid) is required when using the conf option. It is not required when the fasta or gff options are used, but it can be useful with these options, since it can be used to select which sequences JBrowse will import. |
| refids | A comma-delimited list of the database identifiers of sequences to be imported as reference sequences. This option is useful when working with a Chado database that contains data from multiple different species, and those species have at least one chromosome with the same name (e.g. chrX). In this case, the desired chromosome cannot be uniquely identified by name, so it is instead identified by ID. This ID can be found in the 'feature_id' column of 'feature' table in a Chado database. |
Feature Tracks
Feature tracks can be used to visualize localized annotations on a sequence, such as gene models, transcript alignments, SNPs and so forth. JBrowse has several different method of importing annotation data into feature tracks:
- flatfile-to-json.pl - import GFF3 and BED files (recommended for new users)
- biodb-to-json.pl - import from a Bio::DB::SeqFeature::Store database (recommended for users with existing databases)
- bam-to-json.pl - import BAM files
- ucsc-to-json.pl - import UCSC database dumps (.sql and .txt.gz)
Data Formatting
flatfile-to-json.pl
This script inputs a single track into JBrowse. To format multiple tracks for JBrowse, execute the script once for each track.
Terminology: A flat file is a database that exists entirely in a single file. For this script, the flat file must be a GFF3, GFF2, or BED file.
Basic syntax:
bin/flatfile-to-json.pl --[gff|gff2|bed] <flat file> --tracklabel <track name> [options]
Hint: flatfile-to-json.pl simplifies the process of inputting a small number of tracks into JBrowse, since it does not use a config file. If you have many tracks, you will probably want to use a config file, because its structure will make the task of editing tracks easier. In that case, the appropriate script will be biodb-to-json.pl.
| Option | Value |
|---|---|
| gff, gff2, or bed | The name of the file that contains the feature data. The names of these options correspond to the file types, with the exception of gff, which uses a GFF3 file instead of a GFF file. Exactly one of these three options must be used. |
| tracklabel | The internal name that JBrowse will give to this feature track. This option requires a value. |
| key | The external, human-readable label seen on the feature track when it is viewed in JBrowse. The value of key defaults to the value of tracklabel. |
| autocomplete | Dictates what the features of the track will be searchable by after running generate-names.pl. This option can be used with the arguments "label", "alias", "all", or "none". By default, "none" is used.
|
| out | A path to the output directory (default is 'data' in the current directory). |
| cssClass | The css class that will be used to create the feature track. This option makes it possible to choose how the feature track will look by selecting a template class from genome.css. The default css class is 'feature'. |
| getType | Causes the 'type' to be included in the output JSON file. The type is the feature that has been predicted (e.g. promoter site, gene). If a gff file is being used, the type will be in column 3. |
| getPhase | Causes the 'phase' to be included in the output JSON file. The phase describes the reading frame of a DNA (or messenger RNA) sequence. If the phase is relevant, it can have the values 0, 1, or 2; otherwise, the value associated with the phase is '.'. If a gff file is being used, the phase will be in column 8. |
| getSubs | Causes subfeature data to be included in the output JSON file. |
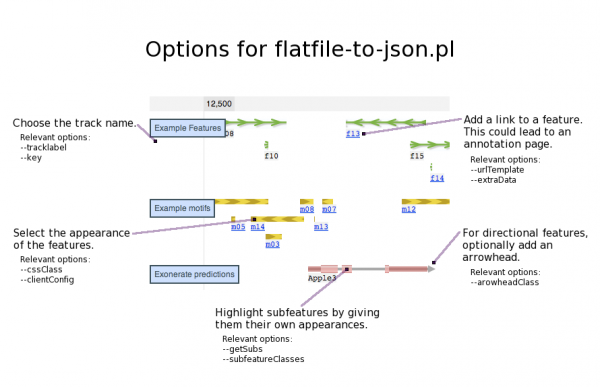
| getLabel | Causes the 'Name' attribute associated with each feature to be included the output JSON file. This will cause a textual name to appear below the features in the track. If a gff3 file is being used, the 'Name' attribute will be in column 9 when it is defined. |
| urlTemplate | A url that your browser will visit when you click on a feature in this track. This is especially useful if you want to link a feature to a page with more information about that feature. |
| arrowheadClass | When this option is used, directional features will be given an arrowhead. The presence and orientation of the arrowhead for each individual feature will depend on data in the input file. Arrowhead classes are defined in genome.css. There is only one that comes with JBrowse (transcript-arrowhead). |
| subfeatureClasses | The css class(es) that will be used for the subfeatures of a feature track. This option makes it possible to choose how the subfeatures will appear. Any of the classes in genome.css can be used for the subfeatures. This option must be used with getSubs in order for subfeatures to appear. |
| clientConfig | Any visual additions or edits for the main features of the track (not for subfeatures). These edits must be specified in JSON syntax. |
| type | The type of feature that will appear in the feature track. This option is useful when the input file contains features of several different types, and you are interested in only having one type of feature (e.g. only having features that are genes) in the feature track. In gff3 files, the type is in the third column. |
| nclChunk | The NCList chunk size. This option should not be used unless an error such as "json or perl structure exceeds maximum nesting level" is encountered. If this error does occur, lower the chunk size (the default is 50000). |
bam-to-json.pl
NOTE: bam-to-json.pl is considered obsolete. While it is still included in JBrowse distributions for the sake of compatibility, its use is not recommended. Most users will want to use the newer BAM Alignment Tracks for direct access to BAM files, with no pre-processing needed.
This script formats a new track into JBrowse based on a BAM file. Tracks added with this script are similar in appearance to tracks added by flatfile-to-json.pl.
Special dependencies: SAMtools, Bio::DB::SAM
Basic syntax:
bin/bam-to-json.pl --bam <bam file> --tracklabel <track name> [options]
| Option | Value |
|---|---|
| bam | The name of the bam file that contains the feature data. This option requires a value. |
| tracklabel | The internal name that JBrowse will give to this feature track. This option requires a value. |
| key | The external, human-readable label seen on the feature track when it is viewed in JBrowse. The value of key defaults to the value of tracklabel. |
| out | A path to the output directory (default is 'data' in the current directory). |
| cssClass | The css class that will be used to create the feature track. This option makes it possible to choose how the feature track will look by selecting a template class from genome.css. The default css class is 'feature'. |
| clientConfig | Any visual additions or edits for the main features of the track (not for subfeatures). These edits must be specified in JSON syntax. |
| nclChunk | The NCList chunk size in bytes. This option should not be used unless an error such as "json or perl structure exceeds maximum nesting level" is encountered. If this error does occur, lower the chunk size (the default is 50000 bytes). |
| compress | This option causes the output JSON files for the track (trackData.json and hist-*.json) to be compressed with gzip. |
biodb-to-json.pl
This script uses a config file to produce a set of feature tracks in JBrowse. It can be used to obtain information from any database with appropriate schema, or from flat files. Because it can produce several feature tracks in a single execution, it is useful for large-scale feature data entry into JBrowse.
Basic syntax:
bin/biodb-to-json.pl --conf <config file> [options]
| Option | Value |
|---|---|
| conf | The name of the JSON configuration file that will be used. This option must be specified. |
| out | A path to the output directory (default is 'data' in the current directory). |
| track | The identifier of a single track that will be updated or added to JBrowse. In the list of key-value pairs comprising an individual track definition in the config file, the identifier will be the value associated with "track". |
| ref | A comma-delimited list of reference sequence names, used to limit database queries to a subset of JBrowse reference sequences. By default, the database is queried for all reference sequences in JBrowse. |
| refid | A comma-delimited list of reference sequence IDs from a Chado database, used to limit database queries to a subset of JBrowse reference sequences. By default, the database is queried for all reference sequences in JBrowse. |
| compress | This option causes the output JSON files for the track (trackData.json and hist-*.json) to be compressed with gzip. |
ucsc-to-json.pl
This script uses data from UCSC genome annotation database. To reach this data, go to hgdownload.cse.ucsc.edu and click the link for the genome of interest. Next, click the "Annotation Database" link. The data relevant to ucsc-to-json.pl (*.sql and *.txt.gz files) can be downloaded from either this page or the FTP server described on this page.
Together, a *.sql and *.txt.gz pair of files (such as cytoBandIdeo.txt.gz and cytoBandIdeo.sql) constitute a database table. Ucsc-to-json.pl uses the *.sql file to get the column labels, and it uses the *.txt.gz file to get the data for each row of the table. For the example pair of files above, the name of the database table is "cytoBandIdeo". This will become the name of the JBrowse track that is produced from the data in the table.
In addition to all of the feature-containing tables that you want to use as JBrowse tracks, you will also need to download the trackDb.sql and trackDb.txt.gz files for the organism of interest.
Basic syntax:
bin/ucsc-to-json.pl --in <directory with files from UCSC> --track <database table name> [options]
Hint: If you're using this approach, it might be convenient to also download the sequence(s) from UCSC. These are usually available from the "Data set by chromosome" link for the particular genome or from the FTP server.
| Option | Value |
|---|---|
| in | A directory containing all of the *.sql and *.txt.gz data from UCSC. This directory must contain the trackDb.sql and trackDb.txt.gz files for the organism of interest, as well as all of the feature-containing tables that you wish to use as JBrowse tracks. |
| track | The name of the database table. If you leave off the .sql or .txt.gz extensions of the table files you wish to use, you will have this value. |
| out | A path to the output directory (default is 'data' in the current directory). |
| cssClass | The css class that will be used to create the feature track. This option makes it possible to choose how the feature track will look by selecting a template class from genome.css. The default css class is 'feature'. |
| arrowheadClass | When this option is used, directional features will be given an arrowhead. The presence and orientation of the arrowhead for each individual feature will depend on data in the input file. Arrowhead classes are defined in genome.css. There is only one that comes with JBrowse (transcript-arrowhead). |
| subfeatureClasses | The css class(es) that will be used for the subfeatures of a feature track. This option makes it possible to choose how the subfeatures will appear. Any of the classes in genome.css can be used for the subfeatures. |
| clientConfig | Any visual additions or edits for the main features of the track (not for subfeatures). These edits must be specified in JSON syntax. |
| nclChunk | The NCList chunk size in bytes. This option should not be used unless an error such as "json or perl structure exceeds maximum nesting level" is encountered. If this error does occur, lower the chunk size (the default is 50000 bytes). |
| compress | This option causes some of the output JSON files (trackData.json and hist-*.json) to be compressed with gzip. |
| sortMem | The maximum amount of RAM (in bytes) to use for sorting the features. The default value is 536870912 bytes (512MiB). |
Configuration Options
The JBrowse track configuration JSON itself has many available options for customization, not all of which are available from the command-line formatting scripts. Below is a comprehensive list of configuration options for HTMLFeatures tracks.
HTMLFeatures Track Options List
| Option | Value |
|---|---|
maxFeatureScreenDensity
|
Maximum density of features to display on the screen. If this is exceeded, will display either feature density histograms (if available), or a "too many features to show" message. The units of this number are features per screen width in pixels. Defaults to 0.5. |
description
|
Comma-separated list of fields in which to look for the description of a feature. Case-insensitive. If set to false or null, no feature description will be shown. Defaults to 'note, description'.
|
maxDescriptionLength
|
Maximum length, in characters, for displayed feature descriptions. |
minSubfeatureWidth
|
Minimum width, in pixels, of the top-level feature for JBrowse to attempt to display its subfeatures. Default 6. |
menuTemplate
|
Optional menu configuration for right-click menus on features. Can be arbitrarily large and complicated. See #Right-click_Context_Menus below. |
hooks→create
|
JavaScript function that creates the parent feature HTML element and returns it. By default this is: function(track,feature) { return document.createElement('div'); }, which creates an HTML div element.
|
hooks→modify
|
JavaScript function that can be used to modify the feature HTML element in any way desired. If set, the function is called with the track object, feature object, and feature HTML element as arguments (signature: function(track, feature, featDiv)).
|
style→featureScale
|
Minimum zoom scale (pixels/basepair) for displaying individual features in the track. Not set by default, and overrides the maxFeatureScreenDensity.
|
style→className
|
CSS class for parent features. Defaults to "feature". |
style→subfeatureClasses
|
Key-value pairs of CSS classes for subfeatures, organized by feature type. Example: { "CDS" : "transcript-CDS","UTR" : "transcript-UTR" } |
style→featureCss
|
Text string of additional CSS rules to add to features. Example: "border-color: purple; background-color: yellow;" |
style→arrowheadClass
|
CSS class of the strand arrowheads to show for this feature. Defaults to 'arrowhead'. If set to null, no arrowhead will be drawn.
|
style→histScale
|
Scale (pixels per basepair) below which the track will attempt to draw feature density histograms instead of features, if available. By default, this is set to 4 times the average feature density (features per basepair) of the track. |
style→labelScale
|
Scale (pixels per basepair) above which feature labels (names) will be shown. By default, this is set to 30 times the average feature density (features per basepair) of the track. |
style→descriptionScale
|
Scale (pixels per basepair) above which long feature descriptions will be shown. By default, this is set to 170 times the average feature density (features per basepair) of the track. |
style→showLabels
|
If set to true, feature labels may be shown. Defaults to true. Set this to false to disable display of feature labels. |
Left-click Behavior
Beginning with JBrowse 1.5.0, the left-clicking behavior of feature tracks is highly configurable. To make something happen when left-clicking features on a track, add an onClick option to the feature track's configuration. In the example configuration below, left-clicks on features will open an embedded popup window showing the results of searching for that feature's name in NCBI's global search, and "search at NCBI" will show in a tooltip when the user hovers over a feature with the mouse:
{ "feature" : [ "mRNA" ], "track" : "ReadingFrame", "category" : "Genes", "class" : "dblhelix", "key" : "Frame usage", "onClick" : { "label": "search at NCBI", "url": "http://www.ncbi.nlm.nih.gov/gquery/?term={name}" } }
For details on all the options supported by onClick, see #Click Configuration Options.
Right-click Context Menus
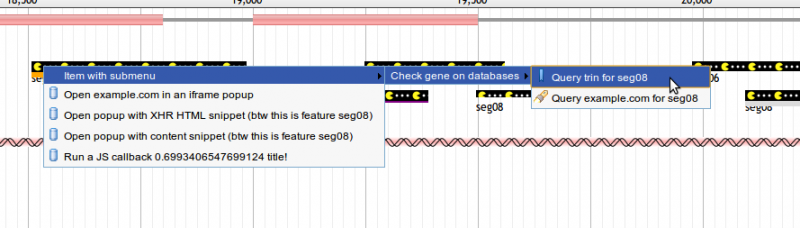
In addition, feature tracks can be configured to display a menu of options when a user right-clicks a menu. Here is an example of a track configured with a multi-level right-click context menu:
{
"feature" : [
"match"
],
"track" : "Alignments",
"category" : "Alignments",
"class" : "feature4",
"key" : "Example alignments",
"hooks": {
"modify": "function( track, feature, div ) { div.style.height = (Math.random()*10+8)+'px'; div.style.backgroundColor = ['green','blue','red','orange','purple'][Math.round(Math.random()*5)];}"
},
"menuTemplate" : [
{
"label" : "Item with submenu",
# hello this is a comment
"children" : [
{
"label" : "Check gene on databases",
"children" : [
{
"label" : "Query trin for {name}",
"iconClass" : "dijitIconBookmark",
"action": "newWindow",
"url" : "http://wiki.trin.org.au/{name}-{start}-{end}"
},
{
"label" : "Query example.com for {name}",
"iconClass" : "dijitIconSearch",
"url" : "http://example.com/{name}-{start}-{end}"
}
]
},
{ "label" : "2nd child of demo" },
{ "label" : "3rd child: this is a track" }
]
},
{
"label" : "Open example.com in an iframe popup",
"title" : "The magnificent example.com (feature {name})",
"iconClass" : "dijitIconDatabase",
"action": "iframeDialog",
"url" : "http://www.example.com?featurename={name}"
},
{
"label" : "Open popup with XHR HTML snippet (btw this is feature {name})",
"title": "function(track,feature,div) { return 'Random XHR HTML '+Math.random()+' title!'; }",
"iconClass" : "dijitIconDatabase",
"action": "xhrDialog",
"url" : "sample_data/test_snippet.html?featurename={name}:{start}-{end}"
},
{
"label" : "Open popup with content snippet (btw this is feature {name})",
"title": "function(track,feature,div) { return 'Random content snippet '+Math.random()+' title!'; }",
"iconClass" : "dijitIconDatabase",
"action": "contentDialog",
"content" : "function(track,feature,div) { return '<h2>{name}</h2><p>This is some test content about feature {name}!</p><p>This message brought to you by the number <span style=\"font-size: 300%\">'+Math.round(Math.random()*100)+'</span>.</p>
},
{
"label" : "function(track,feature,div) { return 'Run a JS callback '+Math.random()+' title!'; }",
"iconClass" : "dijitIconDatabase",
"action": "function( evt ){ alert('Hi there! Ran the callback on feature '+this.feature.get('name')); }"
}
]
}
This configuration results in a context menu like the one pictured below. For details on what each of the options supported by menu items does, see #Click Configuration Options.
Click Configuration Options
A click event can be a string JavaScript callback, like:
"function() { alert('Run any JavaScript you want here!'); }"
Or a structure containing options like:
{ "iconClass" : "dijitIconDatabase", "action" : "iframeDialog", "url" : "http://www.ncbi.nlm.nih.gov/gquery/?term={name}", "label" : "Search for {name} at NCBI", "title" : "function(track,feature,div) { return 'Searching for '+feature.get('name')+' at NCBI'; }" }
The available options for a click action are:
- iconClass
- Used only for click actions in context menus. Usually, you will want to specify one of the Dijit icon classes here. Although they are not well documented, a list of available icon classes can be seen at https://github.com/dojo/dijit/blob/1.7.2/icons/commonIcons.css.
- action
- Either a JavaScript function to run in response to the click (e.g. "function(){..}"), or one of the following special strings: "iframeDialog" - the default - causes the given url to be opened in a popup dialog box within JBrowse, in an
iframeelement. "newWindow" - causes the given url to be opened in a new browser window. "contentDialog" - causes the JavaScript string or callback set in the content option to be displayed in the dialog box. "xhrDialog" - causes the given url to be opened in a popup dialog, containing the HTML fetched from the given url option. The difference between "iframeDialog" and "xhrDialog" is that an iframeDialog's URL should point to a complete web page, while an xhrDialog's URL should point to a URL on the same server (or that supports CORS) that contains just a snippet of HTML (not a complete web page). For those familiar with GBrowse, the xhrDialog is similar to GBrowse popup balloons that use a url:... target, while the contentDialog is similar to a GBrowse popup balloon with a normal target. GBrowse does not have an equivalent of the iframeDialog.
- content
- string (interpolated with feature fields) or JS callback that returns a string. Used only by a contentDialog.
- url
- URL used by newWindow, xhrDialog, or iframeDialog actions.
- label
- descriptive label for the link. In a right-click context menu, this will be the text in the menu item.
- title
- title used for the popup window, or for the mouseover link
Using callbacks to customize feature tracks
JBrowse feature tracks, and individual JBrowse features, can be customized using JavaScript functions you write yourself. These functions are called every time a feature in a track is drawn, and allow you to customize virtually anything about the feature's display. What's more, all of the feature's data is accessible to your customization function, so you can even customize individual features' looks based on their data.
As of JBrowse 1.3.0, feature callbacks are added by directly editing your trackList.json file with a text editor. Unfortunately, due to the limitations of the JSON format currently used for JBrowse configuration, the function must appear as a quoted (and JSON-escaped) string, on a single line. This will be improved in JBrowse 2.0.
Here is an example feature callback, in context in the trackList.json file, that can change a feature's background CSS property (which controls the feature's color) as a function of the feature's name. If the feature's name contains a number that is odd, it give the feature's HTML div element a red background. Otherwise, it gives it a blue background.
{ "autocomplete" : "all", "track" : "ExampleFeatures", "style" : { "className" : "feature2" }, "key" : "Example Features", "feature" : [ "remark" ], "urlTemplate" : "tracks/ExampleFeatures/{refseq}/trackData.json", "hooks": { "modify": "function( track, f, fdiv ) { var nums = f.get('name').match(/\\d+/); if( nums && nums[0] % 2 ) { fdiv.style.background = 'red'; } else { fdiv.style.background = 'blue'; } }" }, "compress" : 0, "label" : "ExampleFeatures", "type" : "FeatureTrack" },
Alignment Tracks (BAM)
JBrowse has several track types that are designed for displaying alignment data, particularly from BAM files.
Alignments2
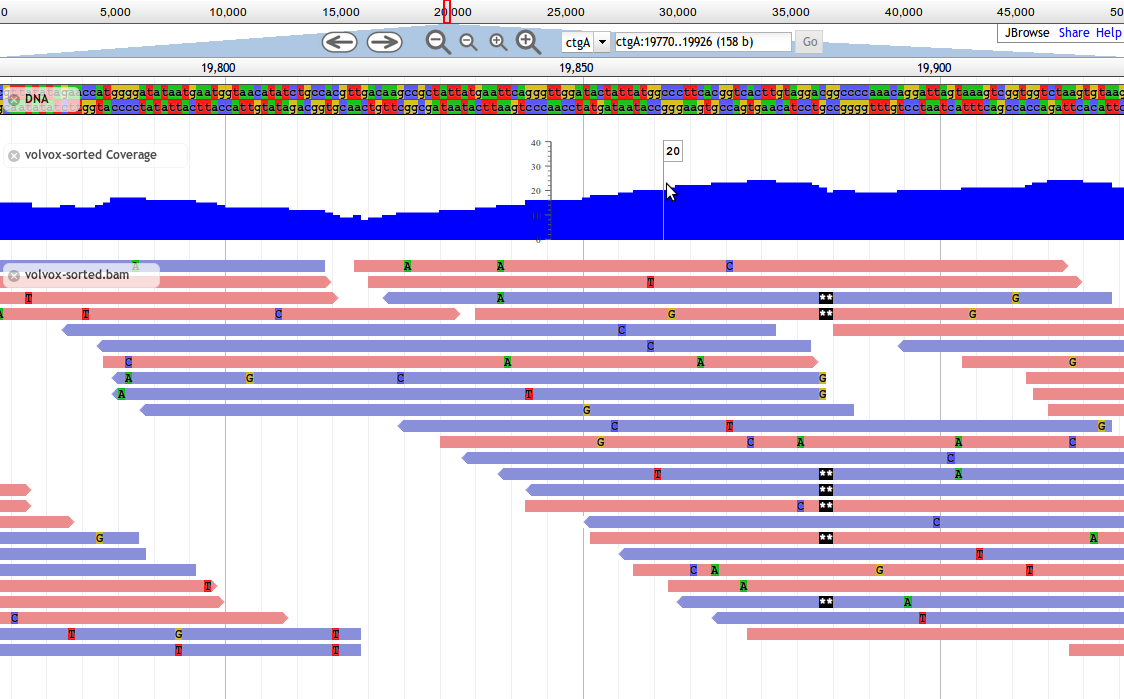
Introduced in JBrowse 1.8.0, Alignments2 tracks are designed to display alignment data, such as from BAM files. This track type shows basepair-level mismatches, insertions, deletions, and skipped regions between aligned reads and the reference, and highlights paired reads in red if their mates are missing.
Base mismatches are displayed based on the contents of the feature's MD field (containing a BAM MD mismatch string), and/or CIGAR field. If your BAM file does not contain MD tags, one common way to generate them is with the samtools calmd command.
Alignments2 is a faster implementation of the older Alignments track type that draws alignments using the HTML5 canvas. In the interest of speed, Alignments2 tracks do not display any text labels or descriptions alongside features, and do not draw arrowheads indicating strandedness, instead relying on color to denote the strand of alignments.
Alignments2 tracks support the same advanced clicking behavior as feature tracks, but do not support right-click menus.
| Option | Value |
|---|---|
maxHeight
|
Available in JBrowse 1.9.0 and later. Maximum displayed height of the track in pixels. Defaults to 1000. Features that would cause the track to grow taller than the maxHeight are not shown. If a section of a track has features that are not drawn because of a maxHeight constraint, a notification is displayed at the bottom of that track section.
|
style→color
|
HTML-style color for drawn alignments. By default, this varies with the alignment's strandedness, and whether its mate pair is missing, using the style→color_fwd_strand, style→color_rev_strand, and style→color_missing_mate variables. To gain complete control over the displayed color, you could set this to be a function callback.
|
style→color_fwd_strand
|
HTML-style color for alignments on the forward strand. Default #EC8B8B, which is a light red. |
style→color_rev_strand
|
HTML-style color for alignments on the reverse strand. Default #898FD8, which is a light blue. |
style→color_missing_mate
|
HTML-style color for alignments with a missing mate. Default #D11919, which is a dark red. |
style→height
|
Height in pixels of each alignment. Default 7. |
style→marginBotton
|
Number of pixels of vertical spacing to put on the bottom of each alignment. Default 1. |
style→showMismatches
|
If true, draw mismatches (SNPs, insertions, deletions, skips) on the alignent. Default true. |
style→mismatchFont
|
CSS string describing the font to use for labeling mismatches. Default "bold 10px Courier New,monospace". |
SNPCoverage
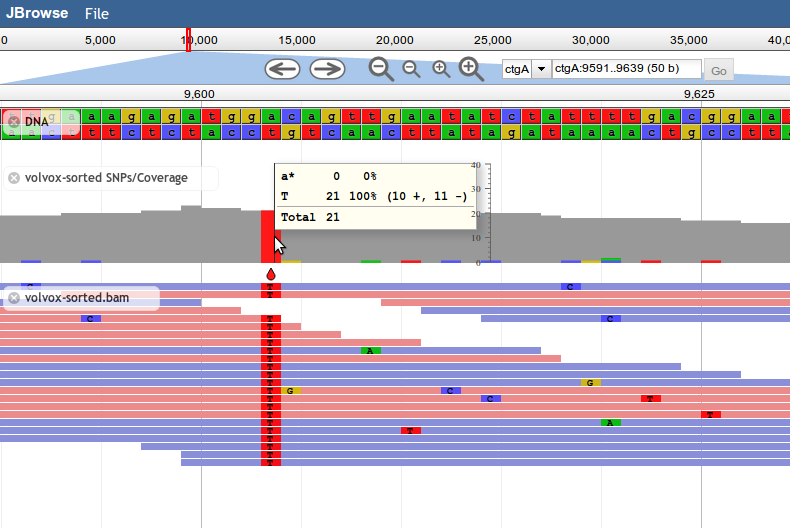
Introduced in JBrowse 1.8.0, SNPCoverage tracks draw the coverage of alignment features along the genome, along with a graphical representation of base-mismatch (possible SNP) distribution, and tables showing frequencies for each mismatching base.
Like the other alignment tracks, base mismatches are displayed based on the contents of the feature's MD field (containing a BAM MD mismatch string).
Note: Since the SNPCoverage track dynamically calculates coverage and putative SNPs directly from alignment data, it is not recommended for use with very dense feature data, such as deep-coverage BAM files. For these types of files, it's recommended to pre-generate a BigWig file of the coverage and a VCF file of putative SNPs, and display those instead.
Example SNPCoverage Configuration
{
"storeClass" : "JBrowse/Store/SeqFeature/BAM",
"urlTemplate" : "../../raw/volvox/volvox-sorted.bam",
"type" : "JBrowse/View/Track/SNPCoverage",
"label" : "volvox-sorted.bam_coverage",
"metadata" : {
"Description" : "SNP/Coverage view of volvox-sorted.bam, simulated resequencing alignments."
},
"key" : "BAM - volvox-sorted SNPs/Coverage"
}
Alignments
Introduced in JBrowse 1.7.0, Alignments tracks are an HTML-based track type for alignment display. They display everything that Alignments2 do, and also can be configured with right-click menus and strand arrowheads.
They display everything that Alignments2 tracks do, plus they support the same configuration options as feature tracks, including advanced clicking behavior, feature modification callbacks, and so forth. The price of this additional capability is that Alignments tracks are significantly slower when used with dense data such as deep BAM alignments.
Alignments2 is recommended over Alignments for most users.
BAM Data Configuration Options
| Option | Value |
|---|---|
urlTemplate
|
URL for the BAM file to display. |
baiUrlTemplate
|
URL for the corresponding BAM index (BAI) file. If not set, this is assumed to be the same URL as urlTemplate with .bai appended.
|
Example BAM Alignments2 track configuration
{ "storeClass" : "JBrowse/Store/SeqFeature/BAM", "urlTemplate" : "../../raw/volvox/volvox-sorted.bam", "label" : "volvox-sorted.bam", "type" : "JBrowse/View/Track/Alignments2", "key" : "volvox-sorted.bam", "style" : { "className" : "alignment", "arrowheadClass" : "arrowhead", "labelScale" : 100 } },
Apache Configuration Note
If you are using the Apache web server, please be aware that the module mime_magic can cause BAM files to be served incorrectly. Some packaged versions of Apache, particularly on Red Hat or CentOS-based systems, are configured with this module turned on by default. We recommend you deactivate this Apache module for the server or directory used to serve JBrowse files.
Wiggle Tracks
Introduced in JBrowse 1.5.0, Wiggle tracks require that the user's browser support HTML <canvas> elements. Currently, the Wiggle tracks are only capable of displaying data from a BigWig files, and BigWig support requires a recent web browser with support for HTML5 typed arrays.
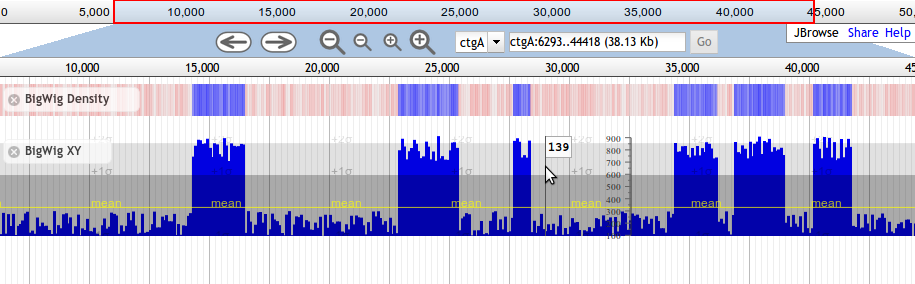
Beginning with JBrowse 1.7.0, there are two available subtypes of Wiggle tracks, Wiggle/XYPlot and Wiggle/Density. The XYPlot wiggle track displays quantitative data as an x/y plot, and the Density displays the data as varying intensities of color.
Example BigWig-based Wiggle XY-Plot Track Configuration
Here is an example track configuration stanza for a Wiggle XY-plot track displaying data directly from a BigWig file. Note that the URL in urlTemplate is relative to the directory where the configuration file is located.
{ "label" : "rnaseq", "key" : "RNA-Seq Coverage" "storeClass" : "JBrowse/Store/SeqFeature/BigWig", "urlTemplate" : "../tests/data/SL2.40_all_rna_seq.v1.bigwig", "type" : "JBrowse/View/Track/Wiggle/XYPlot", "variance_band" : true, "min_score" : -1000, "max_score" : 2000, "style": { "pos_color" : "#FFA600", "neg_color" : "#005EFF", "clip_marker_color" : "red", "height" : 100 } }
Note: numerical values do not appear in quotes.
Example BigWig-based Wiggle Color Density Track Configuration
Here is an example track configuration stanza for a Wiggle color-density track displaying data directly from a BigWig file. It will draw regions with data that is greater than the overall mean as progressively more intense blue, and data that is below the mean as progressively more intense red. Note that the URL in urlTemplate is relative to the directory where the configuration file is located. Also note that green and purple is an ugly color combination. ;-)
{ "label" : "rnaseq", "key" : "RNA-Seq Coverage" "storeClass" : "JBrowse/Store/SeqFeature/BigWig", "urlTemplate" : "my_methylation_data.bw", "type" : "JBrowse/View/Track/Wiggle/Density", "bicolor_pivot" : "mean", "style": { "pos_color": "purple", "neg_color": "green" } }
Wiggle track configuration options
| Option | Value |
|---|---|
origin_color
|
Added in JBrowse 1.7.1, sets the color of the graph origin line in wiggle XY plots. If set to 'none', turns the origin line off. |
bg_color
|
Added in JBrowse 1.7.0, sets the background color for each point at which data is present. Wiggle/Density tracks blend the background color with the pos_color or neg_color to select a color for the region, whereas Wiggle/XYPlot tracks draw the background color on the part of the graph for each data point that is not covered by the pos_color or neg_color. Defaults to transparent for XYPlots, and light gray for Density tracks. |
scale
|
log, default linear Graphing scale, either linear or logarithmic. |
min_score
|
Number. The minimum value to be graphed. Calculated according to autoscale if not provided. |
max_score
|
Number. The maximum value to be graphed. Calculated according to autoscale if not provided. |
autoscale
|
clipped_global|global|z_score If one or more of min_score and max_score options are absent, then these values will be calculated automatically. The autoscale option controls how the calculation is done. A value of local (only available and set as the default in JBrowse 1.9.0 and above) sets the display scale to fit the range of the data being displayed in the current viewing window. A value of global will use global statistics (if available) for the entire quantitative dataset to find min_score and max_score values. z_score will use either ±z_score_bound if it is set, or will use ±4 otherwise. clipped_global is similar to global, except the bounds will be limited to ±z_score_bound, or ±4 if z_score_bound is not set.
|
variance_band
|
1 or 0 If 1, draw a yellow line showing the mean, and two shaded bands showing ±1 and ±2 standard deviations from the mean.
|
z_score_bound
|
for z-score based graphs, the bounds to use. |
data_offset
|
number, default zero. If set, will offset the data display by the given value. For example, a data_offset of -100 would make a data value of 123 be displayed as 23, and a data_offset of 100 would make 123 be displayed as 223.
|
bicolor_pivot
|
"zero"|(num) Where to change from pos_color to neg_color when drawing bicolor plots. Can be "mean", "zero", or a numeric value. Default 0. |
style→pos_color
|
CSS color, default "blue". When drawing bicolor plots, the fill color to use for values that are above the pivot point. Example: |
style→neg_color
|
CSS color, default "red". When drawing bicolor plots, the fill color to use for values that are below the pivot point. |
disable_clip_markers
|
boolean, default false. If true, disables clip markers, which are 2-pixel colored regions at the edge of the graph that indicate when the data value lies outside the displayed range. |
style→clip_marker_color
|
CSS color, defaults to neg_color when in the positive bicolor regime (see bicolor_pivot) and pos_color in the negative bicolor regime. |
style→height
|
Height, in pixels, of the track. Defaults to 100 for XYPlot tracks, and 32 for Density tracks. |
Variant Tracks (VCF)
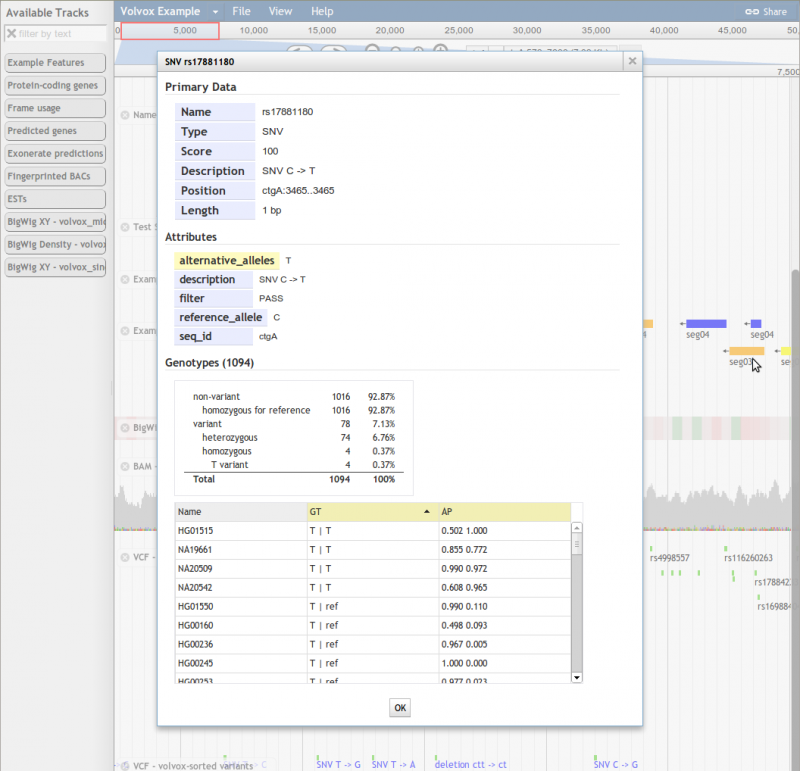
Beginning in JBrowse 1.9.0, JBrowse can display feature data directly from VCF files, and has an HTMLVariants track type that is optimized for displaying the potentially large amounts of detailed data that go with each variant.
Example VCF-based Variant Track Configuration
Here is an example track configuration stanza for a variant track displaying data directly from a VCF file. Note that the URL in urlTemplate is relative to the directory where the configuration file is located.
VCF files used with the VCFTabix must be compressed with bgzip and indexed with tabix, both of which are part of the samtools package.
{ "label" : "mysnps", "key" : "SNPs from VCF", "storeClass" : "JBrowse/Store/SeqFeature/VCFTabix", "urlTemplate" : "../vcf_files/SL2.40_all_rna_seq.v1.vcf.gz", "tbiUrlTemplate": "../vcf_files/SL2.40_all_rna_seq.v1.vcf.gz.tbi", "type" : "JBrowse/View/Track/HTMLVariants" }
Feature Coverage Tracks
Introduced in JBrowse 1.7.0, feature coverage tracks show a dynamically-computed XY-plot of the depth of coverage of features across a genome. One good use of this track type is to provide a
quick coverage plot directly from a BAM file. However, since this track calculates coverage on the fly, it can be very slow when used with large regions or very deep coverage. In this case, it is
recommended to generate a BigWig file containing the coverage data, and display it with a Wiggle/XYPlot or Wiggle/Density track.
Feature coverage tracks are a special type of Wiggle/XYPlot tracks, so the same configuration options apply. There is an additional caveat, however: this kind of track requires the min_score and max_score variables in order to set the Y-axis scaling, since these cannot be quickly determined from raw BAM or other feature data.
Example Feature Coverage Track Configuration for a BAM file
{ "storeClass" : "JBrowse/Store/SeqFeature/BAM", "urlTemplate" : "../../raw/volvox/volvox-sorted.bam", "label" : "volvox-sorted.bam_coverage", "type" : "JBrowse/View/Track/FeatureCoverage", "min_score" : 0, "max_score" : 35, "key" : "volvox-sorted Coverage" }
Image Tracks
In addition to HTML-based feature tracks, JBrowse supports tracks based on pre-generated PNG or JPEG images that are tiled along the reference sequence. Currently, JBrowse ships with two different image track generators: wig-to-json.pl, which generates images showing simple quantitative (wiggle) data, and draw-basepair-track.pl, which draws arcs to show the base pairing structure of RNAs.
wig-to-json.pl
Using a wiggle file, this script creates a single Image track that displays data from the wiggle file. Beginning with JBrowse 1.5, this is no longer the recommended method of displaying wiggle data: it has largely been replaced by the direct-access BigWig data store coupled with the next-generation Wiggle track type. See Wiggle Tracks.
In wiggle data, a numeric value is associated with each nucleotide position in the reference sequence. This is represented in JBrowse as a track that looks like a histogram, where the horizontal axis is for each nucleotide position, and the vertical axis is for the number associated with that position. The vertical axis currently does not have a scale; rather, the heights for each position are relative to each other.
Special dependencies: libpng
In order to use wig-to-json.pl, the code for wig2png must be compiled. Normally, this is done automatically by setup.sh but it can be done manually if necessary. See the Quick Start Tutorial packaged with JBrowse for details.
Basic usage
bin/wig-to-json.pl --wig <wig file> --tracklabel <track name> [options]
Hint: If you are using this type of track to plot a measure of a prediction's quality, where the range of possible quality scores is from some lowerbound to some upperbound (for instance, between 0 and 1), you can specify these bounds with the max and min options.
| Option | Value |
|---|---|
| wig | The name of the wig file that will be used. This option must be specified. |
| tracklabel | The internal name that JBrowse will give to this feature track. This option requires a value. |
| key | The external, human-readable label seen on the feature track when it is viewed in JBrowse. The value of key defaults to the value of tracklabel. |
| out | A path to the output directory (default is 'data' in the current directory). |
| tile | The directory where the tiles, or images corresponding to each zoom level of the track, are stored. Defaults to data/tiles. |
| bgcolor | The color of the track background. Specified as "RED,GREEN,BLUE" in base ten numbers between 0 and 255. Defaults to "255,255,255". |
| fgcolor | The color of the track foreground (i.e. the vertical bars of the wiggle track). Specified as "RED,GREEN,BLUE" in base ten numbers between 0 and 255. Defaults to "105,155,111". |
| width | The width in pixels of each tile. The default value is 2000. |
| height | The height in pixels of each tile. Changing this parameter will cause a corresponding change in the top-to-bottom height of the track in JBrowse. The default value is 100. |
| min | The lowerbound to use for the track. By default, this is the lowest value in the wiggle file. |
| max | The upperbound to use for the track. By default, this will be the highest value in the wiggle file. |
System-specific tips for building wig2png
Apple OS X
You need libpng and a C++ compiler; you can get these with Homebrew, MacPorts, or Fink.
Once you have those installed, you need to compile the JBrowse wiggle-processing program. JBrowse includes a makefile to do this, but you may need to add the paths for libpng and png.h to your compiler's library and include paths. For example, if libpng is in /usr/X11, you can run configure like this:
./configure CXXFLAGS=-I/usr/X11/include LDFLAGS=-L/usr/X11/lib
Depending on which OS X you have (and whether you're using MacPorts/fink/etc.), libpng might be somewhere other than /usr/X11, like /opt/local for example. To find where libpng is, try locate libpng
Ubuntu / Debian Linux
Be sure to install the libpng## and libpng##-dev packages, where ## is a number such as 12 or 15, depending on the version of Ubuntu.
Red Hat/CentOS/Fedora Linux
Be sure to install the libpng and libpng-devel packages.
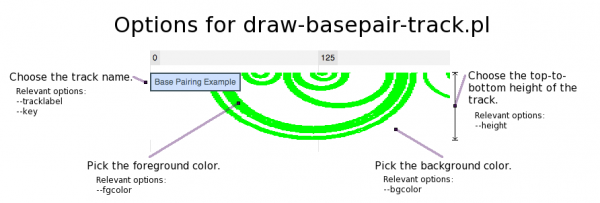
draw-basepair-track.pl
This script inputs a single base pairing track into JBrowse. A base pairing track is a distinctive track type that represents base pairing between nucleotides as arcs. In addition, it is intended to demonstrate the Perl API for writing your own image track generators.
Basic usage
bin/draw-basepair-track.pl --gff <gff file> --tracklabel <track name> [options]
| Option | Value |
|---|---|
| gff | The name of the gff file that will be used. This option must be specified. |
| tracklabel | The internal name that JBrowse will give to this feature track. This option requires a value. |
| key | The external, human-readable label seen on the feature track when it is viewed in JBrowse. The value of key defaults to the value of tracklabel. |
| out | A path to the output directory (default is 'data' in the current directory). |
| tile | The directory where the tiles, or images corresponding to each zoom level of the track, are stored. Defaults to data/tiles. |
| bgcolor | The color of the track background. Specified as "RED,GREEN,BLUE" in base ten numbers between 0 and 255. Defaults to "255,255,255". |
| fgcolor | The color of the track foreground (i.e. the base pairing arcs). Specified as "RED,GREEN,BLUE" in base ten numbers between 0 and 255. Defaults to "0,255,0". |
| width | The width in pixels of each tile. The default value is 2000. |
| height | The height in pixels of each tile. Changing this parameter will cause a corresponding change in the top-to-bottom height of the track in JBrowse. The default value is 100. |
| thickness | The thickness of the base pairing arcs in the track. The default value is 2. |
| nolinks | Disables use of file system links to compress duplicate image files. |
Name Searching and Autocompletion
The JBrowse search box auto-completes the names of features and reference sequences that are typed into it. After loading all feature and reference sequence data into a JBrowse instance (with prepare-refseqs.pl, flatfile-to-json.pl, etc.), generate-names.pl must be run to build the indexes used for name searching and autocompletion.
Autocompletion Configuration
Several settings are available to customize the behavior of autocompletion. Most users will not need to configure any of these variables.
| Option | Value |
|---|---|
autocomplete→stopPrefixes
|
Array of word-prefixes for which autocomplete will be disabled. For example, a value of ['foo'] will prevent autocompletion when the user as typed 'f', 'fo', or 'foo', but autocompletion will resume when the user types any additional characters.
|
autocomplete→resultLimit
|
Maximum number of autocompletion results to display. Defaults to 15. |
autocomplete→tooManyMatchesMessage
|
Message displayed in the autocompletion dropdown when more than autocomplete→resultLimit matches are displayed. Defaults to 'too many matches to display'.
|
generate-names.pl
This script builds indexes of features by label (the visible name below a feature in JBrowse) and/or by alias (a secondary name that is not visible in the web browser, but may be present in the JSON used by JBrowse).
To search for a term, type it in the autocompleting text box at the top of the JBrowse window.
Basic syntax:
bin/generate-names.pl [options]
Note that generate-names.pl does not require any arguments. However, some options are available:
| Option | Value |
|---|---|
| out | A path to the output directory (default is 'data/' in the current directory). |
| thresh | A lower-bound on the Patricia trie chunk size. Specifically, the lowest possible chunk size is (thresh + 1). The default value is 200. In this context, a chunk is a group of connected Patricia trie nodes that can be visualized as a single entity, and the chunk size is the total number of genomic features contained in a chunk. The lower the value of thresh, the more chunks there will be. |
| verbose | This setting causes information about the division of nodes into chunks to be printed to the screen. |
Removing Tracks
JBrowse has a script to remove individual tracks: remove-track.pl. Run it with the --help option to see a comprehensive usage message:
bin/remove-track.pl --help
Compressing data on the server
Starting with JBrowse 1.3.0, server-side data-formatting scripts support a
--compress option to compress (gzip) feature and sequence
data to conserve server disk space and reduce server CPU load even
further. Using this option requires some additional web server configuration.
- For Apache
-
AllowOverride FileInfo(orAllowOverride All) must be set for the JBrowse data directories in order to use the.htaccessfiles generated by the formatting scripts. -
mod_headersmust be installed and enabled, and if the web server is usingmod_gzipormod_deflate,mod_setenvifmust also be installed and enabled.
- For nginx
- A configuration snippet like the following should be included in the configuration:
location ~* "\.(json|txt)z$" {
add_header Content-Encoding gzip;
gzip off;
types { application/json jsonz; }
}
Controlling JBrowse with the URL Query String
As shipped, JBrowse provides a number of options for changing the current view in the browser by adding query parameters to the URL. Developers, please note that these URL parameters are handled only in the default index.html that ships with JBrowse; the JBrowse code itself does not pay attention to URL parameters.
Basic syntax:
http://<server>/<path to jbrowse>?loc=<location string>&tracks=<tracks to show>
data
Optional relative (or absolute) URL of the base directory from which JBrowse will load data. Defaults to 'data'.
loc
The initial genomic position which will be visible in the viewing field. Possible input strings are:
"Chromosome"+":"+ start point + ".." + end point
A chromosome name/ID followed by “:”, starting position, “..” and end position of the genome to be viewed in the browser is used as an input. Chromosome ID can be either a string or a mix of string and numbers. “CHR” to indicate chromosome may or may not be used. Strings are not case-sensitive. If the chromosome ID is found in the database reference sequence (RefSeq), the chromosome will be shown from the starting position to the end position given in URL.
example) ctgA:100..200
Chromosome ctgA will be displayed from position 100 to 200.
OR start point + ".." + end point
A string of numerical value, “..” and another numerical value is given with the loc option. JBrowse navigates through the currently selected chromosome from the first numerical value, start point, to the second numerical value, end point.
example) 200..600
OR center base
If only one numerical value is given as an input, JBrowse treats the input as the center position. Then an arbitrary region of the currently selected gene is displayed in the viewing field with the given input position as basepair position on which to center the view.
OR feature name/ID
If a string or a mix of string and numbers are entered as an input, JBrowser treats the input as a feature name/ID of a gene. If the ID exists in the database RefSeq, JBrowser displays an arbitrary region of the feature from the the position 0, starting position of the gene, to a certain end point.
example) ctgA
tracks
parameters are comma-delimited strings containing track names, each of which should correspond to the "label" element of the track information dictionaries that are currently viewed in the viewing field. Names for the tracks can be found in data/trackInfo.js in jbrowse-1.2.1 folder.
example) DNA,knownGene,ccdsGene,snp131,pgWatson,simpleRepeat
Embedded mode
JBrowse's included index.html file supports three URL query arguments that can turn off the JBrowse track list, navigation bar, and overview bar, respectively. When all three of these are turned off, the only thing visible are the displayed tracks themselves, and JBrowse could be said to be running in a kind of "embedded mode".
The three parameters used for this are nav, tracklist, and overview. If any of these are set to 0, that part of the JBrowse interface is hidden.
For example, you could put the embedded-mode JBrowse in an iframe, like this:
<html>
<head>
<title>JBrowse Embedded</title>
</head>
<body>
<h1>Embedded Volvox JBrowse</h1>
<div style="width: 400px; margin: 0 auto;">
<iframe style="border: 1px solid black" src="../../index.html?data=sample_data/json/volvox&tracklist=0&nav=0&overview=0&tracks=DNA%2CExampleFeatures%2CNameTest%2CMotifs%2CAlignments%2CGenes%2CReadingFrame%2CCDS%2CTranscript%2CClones%2CEST" width="300" height="300">
</iframe>
</div>
</body>
</html>
Data Export
Starting with version 1.7.0, JBrowse users can export track data in a variety of formats for either specific regions, or for entire reference sequences. Export functionality can also be limited and disabled on a per-track basis using the configuration variables listed below.
Data Formats
Current supported export data formats are:
- FASTA (sequence tracks)
- GFF3 (all tracks)
- bed (feature and alignment tracks)
- bedGraph (wiggle tracks)
- Wiggle (wiggle tracks)
Export Configuration
Each track in JBrowse that can export data supports the following configuration variables.
| Option | Value |
|---|---|
noExport
|
If true, disable all data export functionality for this track. Default false. |
maxExportSpan
|
Maximum size of the a region, in bp, that can be exported from this track. Default 500 Kb. |
maxExportFeatures
|
Maximum number of features that can be exported from this track. Not set by default. |
Track Metadata
JBrowse supports two ways to add data that describes tracks (track metadata): it can either be embedded directly in the track's configuration stanza, or it can come from a separate file that JBrowse loads. Track metadata is shown both when a user clicks "About this track" from a track's menu, and in the faceted track selector if it is in use.
Embedding Track Metadata in JBrowse Configuration
Each track configuration stanza can include a metadata item that contains items of data that describe the track. For example, to describe a BAM track containing alignments of RNA-seq reads from Volvox carteri under conditions of caffeine starvation, a track configuration might contain:
{ "storeClass" : "JBrowse/Store/SeqFeature/BAM", "urlTemplate" : "../../raw/volvox/volvox-sorted.bam", "style" : { "className" : "alignment", "arrowheadClass" : "arrowhead", "labelScale" : 100 }, "label" : "volvox-sorted.bam", "type" : "JBrowse/View/Track/Alignments", "key" : "volvox-sorted.bam", "metadata": { "Description": "RNA-seq", "Conditions": "caffeine starvation", "Species": "Volvox carteri", "Data Provider": "Robert Buels Institute for Example Data" } }
Loading Track Metadata from Files
To add track metadata from an external file to JBrowse, add a trackMetadata section to the JBrowse configuration.
JBrowse currently supports track metadata that in Excel-compatible comma-separated-value (CSV) format, but additional track metadata backends are relatively easy to add. Write the JBrowse mailing list if you have a strong need to use another format.
| Option | Value |
|---|---|
trackMetadata.sources
|
Array of source definitions, each of which takes the form { type: 'csv', url: '/path/to/file' }. The url is interpreted as relative to the url of the page containing JBrowse (index.html in default installations). Source definitions can also contain a class to explicitly specify the JavaScript backend used to handle this source.
|
trackMetadata.indexFacets
|
Optional array of facet names that should be the only ones made searchable. This can be used improve the speed and memory footprint of JBrowse on the client by not indexing unused metadata facets. |
Example
Configuration:
trackMetadata: { indexFacets: [ 'category','organism','target','technique','principal_investigator', 'factor','developmental-stage','strain','cell-line','tissue','compound', 'temperature' ], sources: [ { type: 'csv', url: 'myTrackMetaData.csv' } ] }
Track metadata CSV:
| label | technique | factor | target | principal_investigator | submission | category | type | Developmental-Stage |
|---|---|---|---|---|---|---|---|---|
| fly/White_INSULATORS_WIG/BEAF32 | ChIP-chip | BEAF-32 | Non TF Chromatin binding factor | White, K. | 21 | Other chromatin binding sites | data set | Embryos 0-12 hr |
| fly/White_INSULATORS_WIG/CP190 | ChIP-chip | CP190 | Non TF Chromatin binding factor | White, K. | 22 | Other chromatin binding sites | data set | Embryos 0-12 hr |
| fly/White_INSULATORS_WIG/GAF | ChIP-chip | GAF | Non TF Chromatin binding factor | White, K. | 23 | Other chromatin binding sites | data set | Embryos 0-12 hr |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
Note that the label for each track metadata row must correspond to the label in the track configuration for the track it describes.
Faceted Track Selector
Starting with version 1.4.0, JBrowse has an optional "faceted" track selector designed for sites with hundreds or even thousands of tracks in a single JBrowse instance. This track selector allows users to interactively search for the tracks they are interested in based on the metadata for each track.
An example of a faceted track selector in action with about 1,800 tracks can be seen here. This is an example installation containing a snapshot of modENCODE track metadata. Note that the track data and reference sequences in this example are not real (they are actually all just copies of the same volvox test track), this is just an example of the faceted track selector in action.
The Faceted track selector takes all sources of track metadata, aggregates them, and makes the tracks searchable using this metadata. By default, tracks only have a few default metadata facets that come from the track configuration itself. After initially turning on the faceted track selector, most users will want to add their own metadata for the tracks: see #Defining Track Metadata below. To enable the faceted track selector in the JBrowse configuration, set trackSelector→type to Faceted.
There are some other configuration variables that can be used to customize the display of the track selector. Most users will want to set both of these variables to customize the columns and facets shown in the track selector.
| Option | Value |
|---|---|
trackSelector→displayColumns
|
Array of which facets should be displayed as columns in the track list. Columns are displayed in the order given. If not provided, all facets will be displayed as columns, in lexical order. |
trackSelector→renameFacets
|
Object containing "display names" for some or all of the facet names. For example, setting this to { 'developmental-stage': 'Conditions' } would display "Conditions" as the name of the developmental-stage facet.
|
trackSelector→escapeHTMLInData
|
Beginning in JBrowse 1.6.5, if this is set to true or 1 prevents HTML code that may be present in the track metadata from being rendered. Instead, the HTML code itself will be shown.
|
trackSelector→selectableFacets
|
Optional array of which facets should be displayed as facet selectors. The selectors for these appear in the order in which they are specified here. |
Example Faceted Track Selector Configuration
trackSelector: { type: 'Faceted', displayColumns: ['key', 'organism', 'technique', 'target', 'factor', 'developmental-stage','principal_investigator','submission' ], renameFacets: { 'developmental-stage': 'Conditions', submission: 'Submission ID' }, selectableFacets: ['organism','technique','developmental-stage','factor'] }
Dataset Selector
Beginning in version 1.9.0, JBrowse supports an optional dropdown dataset selector that appears in the menu bar in the upper left corner of the JBrowse display. The selector displays a list of dataset names based on the datasets key in the JBrowse configuration. When the selector is changed, a page reload is triggered, with the browser navigating to the URL associated with the selected dataset.
To set which value of the dataset selector is currently selected, the configuration variable dataset_id should be set to the name of the corresponding entry in the datasets list.
The selector is only shown if both datasets and dataset_id are set in the configuration.
Example Dataset Switching Configuration
- in
jbrowse_conf.json
datasets: {
volvox: { url: '?data=sample_data/json/volvox', name: 'Volvox Example' },
modencode: { url: '?data=sample_data/json/modencode', name: 'MODEncode Example' },
yeast: { url: '?data=sample_data/json/yeast', name: 'Yeast Example' }
}
- in
sample_data/json/volvox/trackList.json
dataset_id: 'volvox'
- in
sample_data/json/modencode/trackList.json
dataset_id: 'modencode'
- in
sample_data/json/yeast/trackList.json
dataset_id: 'yeast'
Anonymous Usage Statistics
JBrowse instances report usage statistics to the JBrowse
developers. This data is very important to the JBrowse project,
since it is used to make the case to grant agencies for continuing
to fund JBrowse development. No research data is transmitted, the
data collected is limited to standard Google Analytics, along with
a count of how many tracks the JBrowse instance has, how many
reference sequences are present, their average length, and what
types of tracks (wiggle, feature, etc) are present. Users can
disable usage statistics by setting suppressUsageStatistics: true
in the JBrowse configuration.
Advanced Topics
Data Format Specification: LazyNCList Feature Store
JBrowse uses lazily-loaded nested containment lists (LazyNCLists) as an efficient format for storing feature data in pre-generated static files. A nested containment list is a tree data structure in which the nodes of the tree are intervals themselves features, and edges connecting features that lie `within the bounds of (but are not subfeatures of) another feature. It has some similarities to an R tree. For more on NClists, see the Alekseyenko paper.
This data format is currently used in JBrowse 1.3 for tracks of type FeatureTrack, and the code that actually reads this format is in SeqFeatureStore/NCList.js and ArrayRepr.js.
The LazyNCList format can be broken down into two distinct subformats: the LazyNCList itself, and the array-based JSON representation of the features themselves.
Array Representation (ArrayRepr)
For speed and memory efficiency, JBrowse feature JSON represents features as arrays instead of objects. This is because the JSON representation is much more compact (saving a lot of disk space), and many browsers significantly optimize JavaScript Array objects over more general objects.
Each feature is represented as an array of the form [ class, data, data, ... ], where the class is an integer index into the store's classes array (more on that in the next section). Each of the elements in the classes array is an array representation that defines the meaning of each of the the elements in the feature array.
An array representation specification is encoded in JSON as (comments added):
{
"attributes": [ // array of attribute names for this representation
"AttributeNameForIndex1",
"AttributeNameForIndex2",
...
],
"isArrayAttr": { // list of which attributes are themselves arrays
"AttributeNameForIndexN": 1,
...
}
}
Lazy Nested-Containment Lists (LazyNCList)
A JBrowse LazyNCList is a nested containment list tree structure stored as one JSON file that contains the root node of the tree, plus zero or more "lazy" JSON files that contain subtrees of the main tree. These subtree files are lazily fetched: that is, they are only fetched by JBrowse when they are needed to display a certain genomic region.
On disk, the files in an LazyNCList feature store look like this:
# stats, metadata, and nclist root node data/tracks/<track_label>/<refseq_name>/trackData.json # lazily-loaded nclist subtrees data/tracks/<track_label>/<refseq_name>/lf-<chunk_number>.json # precalculated feature densities data/tracks/<track_label>/<refseq_name>/hist-<bin_size>.json ...
Where the trackData.json file is formatted as (comments added):
{
"featureCount" : 4293, // total number of features in this store
"histograms" : { // information about precalculated feature-frequency histograms
"meta" : [
{ // description of each available bin-size for precalculated feature frequencies
"basesPerBin" : "100000",
"arrayParams" : {
"length" : 904,
"chunkSize" : 10000,
"urlTemplate" : "hist-100000-{Chunk}.json"
}
},
... // and so on for each bin size
],
"stats" : [
{ // stats about each precalculated set of binned feature frequencies
"basesPerBin" : "100000", // bin size in bp
"max" : 51, // max features per bin
"mean" : 4.93030973451327 // mean features per bin
},
...
]
},
"intervals" : {
"classes" : [ // classes: array representations used in this feature data (see ArrayRepr section above)
{
"isArrayAttr" : {
"Subfeatures" : 1
},
"attributes" : [
"Start",
"End",
"Strand",
"Source",
"Phase",
"Type",
"Id",
"Name",
"Subfeatures"
]
},
...
{ // the last arrayrepr class is the "lazyClass": fake features that point to other files
"isArrayAttr" : {
"Sublist" : 1
},
"attributes" : [
"Start",
"End",
"Chunk"
]
}
],
"nclist" : [
[
2, // arrayrepr class 2
12962, // "Start" minimum coord of features in this subtree
221730, // "End" maximum coord of features in this subtree
1 // "Chunk" (indicates this subtree is in lf-1.json)
],
[
2, // arrayrepr class 2
220579, // "Start" minimum coord of features in this subtree
454457, // "End" maximum coord of features in this subtree
2 // "Chunk" (indicates this subtree is in lf-2.json)
],
...
],
"lazyClass" : 2, // index of arrayrepr class that points to a subtree
"maxEnd" : 90303842, // maximum coordinate of features in this store
"urlTemplate" : "lf-{Chunk}.json", // format for lazily-fetched subtree files
"minStart" : 12962 // minimum coordinate of features in this store
},
"formatVersion" : 1
}
Data Format Specification: Fixed-Resolution Tiled Image Store
JBrowse can display tracks composed of precalculated image tiles, stretching the tile images horizontally when necessary. The JBrowse Volvox example data has a wiggle data track that is converted to image tiles using the included wig2png program, but any sort of image tiles can be displayed if they are laid out in this format.
The files for a tiled image track are structured by default like this:
data/tracks/<track_label>/<refseq_name>/trackData.json data/tracks/<track_label>/<refseq_name>/<zoom_level_urlPrefix>/<index>.png ... (and so on, for many more PNG image files)
Where the PNG files are the image tiles themselves, and trackData.json contains metadata about the track in JSON format, including available zoom levels, the width and height of the image tiles, their base resolution (number of reference sequence base pairs per image tile), and statistics about the data (such as the global minimum and maximum of wiggle data).
The structure of the trackData.json file is:
{
"tileWidth": 2000, // width of all image tiles, in pixels
"stats" : { // any statistics about the data being represented
"global_min": 100,
"global_max": 899
},
"zoomLevels" : [ // array describing what resolution levels are available
{ // in the precalculated image tiles
"urlPrefix" : "1/",
"height" : 100,
"basesPerTile" : 2000
},
... (and so on, for zoom levels in order of decreasing resolution / increasing bases per tile )
]
}
To see a working example of this in action, see the contents of sample_data/json/volvox/tracks/volvox_microarray.wig/ctgA after the Volvox wiggle sample data has been formatted.
The code for working with this tiled image format in JBrowse 1.3 is in TiledImageStore/Fixed.js.
Other Global Configuration
JBrowse supports some other configuration variables that can change the overall behavior of the browser.
| Option | Description |
|---|---|
locationBoxLength
|
The desired size, in characters of the location search box. If not set, the size of the location box is calculated to fit the largest location string that is likely to be produced, based on the length of the reference sequences and the length of their names. Added in JBrowse 1.7.0. |
css
|
Used to add additional CSS code to the browser at runtime. Can be an array of either strings containing CSS statements, or URLs for CSS stylesheets to load (as {url: "/path/to/my.css"}). CSS can of course also be added outside of JBrowse, at the level of the HTML page where JBrowse runs. Added in JBrowse 1.6.2.
|
theme
|
Allows changing the graphical theme from the default Dijit "tundra" theme. Added in JBrowse 1.7.0. |
Using JBrowse with Existing Databases
Extract data and reformat
The JBrowse formatting tools biodb-to-json.pl and prepare-refseq.pl can extract data from existing databases that are supported by BioPerl Bio::DB::* adapters, such as GBrowse databases created by bp_seqfeature_load.pl, or Chado databases. Both tools accepts a configuration file in JSON format that contains the details about how to connect to a given database, and which data to extract from it, and which JBrowse feature tracks to create to display the data.
For example, to extract data from a Chado schema in PostgreSQL, one might start with a configuration like:
{ "description": "D. melanogaster (release 5.37)", "db_adaptor": "Bio::DB::Das::Chado", "db_args": { "-dsn": "dbi:Pg:dbname=fruitfly;host=localhost;port=5432", "-user": "yourusername", "-pass": "yourpassword" }, ... }
In the database source name (dsn) argument, 'dbi:Pg' indicates that you are using PostgreSQL, and the dbname, host, and port were specified when the database was created with PostgreSQL's createdb command. The user and pass arguments were specified when the PostgreSQL user account was created with the createuser command. Collectively, these arguments identify the database and give the Bio::DB::Das::Chado object access to it. Other adaptors (Bio::DB::SeqFeature::Store, Bio:DB::GFF, etc.) will require similar information.
Example Configuration
Here is a sample configuration file, usable with biodb-to-json.pl and prepare-refseqs.pl, with each line explained. Note that, in order for this config file to work, it would be necessary to remove the grey comments (since JSON does not support them). Also, notice that the config file is divided into two parts, a header section that contains information about the database, and a body section that contains information about the feature tracks.
{
This is the header. It contains information about the database.
description: a brief textual description of the data source.
"description": "D. melanogaster (release 5.37)",
db_adaptor: a perl module with methods for opening databases and extracting
information. This will normally be either Bio::DB::SeqFeature::Store,
Bio::DB::Das::Chado, or Bio::DB::GFF, but it can also be the name of any
other perl module that implements the Bio::SeqFeatureI interface.
"db_adaptor": "Bio::DB::SeqFeature::Store",
db_args: arguments required to produce an instance of the db_adaptor. The
required arguments can be found by searching for the db_adaptor on the CPAN
website.
"db_args": {
adaptor: With Bio::DB::SeqFeature::Store, a value of "memory"
for the adaptor indicates that the data is stored somewhere in
the file system. Alternatively, it might have been stored in a
database such as MySQL or BerkeleyDB.
"-adaptor": "memory",
dir: given the "memory" argument for the adaptor, this is the
file system path to the location in memory where the data is
stored. Data will automatically be extracted from any *.gff
or *.gff3 files in this directory.
"-dir": "/Users/stephen/Downloads/dmel_r5.37"
},
This is the body. It contains information about the feature tracks.
TRACK DEFAULTS: The default options for every track.
"TRACK DEFAULTS": {
class: same as 'cssClass' in flatfile-to-json.pl.
"class": "feature"
},
tracks: information about each individual track.
"tracks": [
Information about the first track.
{
track: same as 'tracklabel' in flatfile-to-json.pl.
"track": "gene",
key: same meaning as in flatfile-to-json.pl.
"key": "Gene Span",
feature: an array of the feature types that will be used for the track.
Similar to 'type' in flatfile-to-json.pl.
"feature": ["gene"],
autocomplete: same meaning as in flatfile-to-json.pl.
"autocomplete": "all",
"class": "feature2",
urlTemplate: same meaning as in flatfile-to-json.pl. Note how
urlTemplate is being used with a variable called "feature_id" defined
in extraData. In this way, different features in the same track can
be linked to different pages on FlyBase.
"urlTemplate": "http://flybase.org/cgi-bin/fbidq.html?{feature_id}",
extraData: same as in flatfile-to-json.pl.
"extraData": {"feature_id": "sub {shift->attributes(\"load_id\");}"}
},
Information about the second track.
{
"track": "mRNA",
"feature": ["mRNA"],
"autocomplete": "alias",
subfeatures: similar to 'getSubs' in flatfile-to-json.pl.
"subfeatures": true,
"key": "mRNA",
"class": "transcript",
subfeature_classes: same as 'subfeatureClasses' in flatfile-to-json.pl.
"subfeature_classes": {
"CDS": "transcript-CDS",
"five_prime_UTR": "transcript-five_prime_UTR",
"three_prime_UTR": "transcript-three_prime_UTR"
},
arrowheadClass: same meaning as in flatfile-to-json.pl.
"arrowheadClass": "transcript-arrowhead",
clientConfig: same meaning as in flatfile-to-json.pl.
"clientConfig": {
"histScale":5
},
"urlTemplate": "http://flybase.org/cgi-bin/fbidq.html?{feature_id}",
"extraData": {"feature_id": "sub {shift->attributes(\"load_id\");}"}
}
]
}
Using JBrowse with Existing Web Services
Users can extend JBrowse's functionality to with their own JavaScript code using the JBrowse plugin system. For an overview of plugins and their structure, see #Writing JBrowse Plugins.
To use JBrowse with an existing set of web services, users will want to implement a JBrowse Store module in JavaScript that can fetch data from them and convert it into the internal JavaScript object representations that JBrowse expects. In general terms, the steps to follow to do this would be:
1. Create a new plugin using bin/new-plugin.pl or manually.
1. Enable the plugin by adding its name to plugins in the JBrowse configuration (in jbrowse_conf.json, in the constructor arguments in index.html, or elsewhere).
1. Create a new data store class in the plugin's JS directory that inherits from JBrowse/Store/SeqFeature and overrides its methods.
Example custom JBrowse store class
In plugins/MyPlugin/js/Store/SeqFeature/FooBaseWebServices.js, usable in store or track configurations as MyPlugin/Store/SeqFeature/FooBaseWebServices.
/**
* Example store class that uses Dojo's XHR libraries to fetch data
* from backend web services. In the case of feature data, converts
* the data into JBrowse SimpleFeature objects (see
* JBrowse/Model/SimpleFeature.js) but any objects that support the
* same methods as SimpleFeature are fine.
*/
define([
'dojo/_base/declare',
'dojo/_base/array',
'dojo/request/xhr',
'JBrowse/Store/SeqFeature',
'JBrowse/Model/SimpleFeature'
],
function( declare, array, xhr, SeqFeatureStore, SimpleFeature ) {
return declare( SeqFeatureStore, {
constructor: function( args ) {
// perform any steps to initialize your new store.
},
getGlobalStats: function( statsCallback, errorCallback ) {
var thisB = this;
xhr.get( this.config.baseUrl+'my/webservice/url',
{ handleAs: 'json' }
).then(
function( data ) {
data = thisB._transformStatsDataSomehow( data );
statsCallback( data );
},
errorCallback
);
},
getRegionStats: function( query, statsCallback, errorCallback ) {
var thisB = this;
xhr.get( this.config.baseUrl+'my/other/webservice/url',
{ handleAs: 'json', query: query }
).then(
function( data ) {
data = thisB._transformStatsDataSomehow( data );
statsCallback( data );
},
errorCallback
);
},
getFeatures: function( query, featureCallback, finishCallback, errorCallback ) {
var thisB = this;
xhr.get( this.config.baseUrl+'my/features/webservice/url',
{ handleAs: 'json', query: query }
).then(
function( featuredata ) {
// transform the feature data into feature
// objects and call featureCallback for each
// one. for example, the default REST
// store does something like:
array.forEach( featuredata || [],
function( featureKeyValue ) {
var feature = new SimpleFeature({
data: featureKeyValue
});
featureCallback( feature );
});
// call the endCallback when all the features
// have been processed
endCallback();
},
errorCallback
);
}
});
});
Writing JBrowse-compatible Web Services
Beginning in version 1.9.0, JBrowse ships with a REST data store adapter (JBrowse/Store/SeqFeature/REST) that can provide feature, sequence, and quantitative data for display by any of JBrowse's track types. To use it, a developer can implement server-side web services that satisfy the REST API it expects.
JBrowse REST Feature Store API
The JBrowse REST feature store requires the following server resources.
GET (base)/stats/global
Required. Returns a JSON object containing global statistics about the features served by this store.
Example:
{ "featureDensity": 0.02, "featureCount": 234235, "scoreMin": 87, "scoreMax": 87, "scoreMean": 42, "scoreStdDev": 2.1 }
None of the attributes in the example above are required to be present. However, if the store is primarily providing positional data (such as genes), it is recommended to provide at least featureDensity (average number of features per basepair), since JBrowse uses this metric to make many decisions about how to display features. For stores that primarily provide quantitative data, it is recommended to also provide score statistics.
GET (base)/stats/region/(refseq_name)?start=123&end=456
Optional, but recommended. Get statistics for a particular region. Returns the same format as stats/global above, but with statistics that apply only to the region specified.
The refseq name URL component, and the start and end query parameters specify the region of interest. Statistics should be calculated for all features that intersect the region in question. start and end are in interbase coordinates.
NOTE: If this is not implemented, the statistics will be calculated as needed by actually fetching feature data for the region in question.
GET (base)/features/(refseq_name)?start=123&end=456
Required. Fetch feature data (including quantitative data) for the specified region.
The refseq name URL component, and the start and end query parameters specify the region of interest. Statistics should be calculated for all features that intersect the region in question. start and end are in interbase coordinates.
Example return JSON:
{ "features": [ /* minimal required data */ { "start": 123, "end": 456 }, /* typical quantitative data */ { "start": 123, "end": 456, "score": 42 }, /* typical processed transcript with subfeatures */ { "type": "mRNA", "start": 5975, "end": 9744, "score": 0.84, "strand": 1, "name": "au9.g1002.t1", "subfeatures": [ { "type": "five_prime_UTR", "start": 5975, "end": 6109, "score": 0.98, "strand": 1 }, { "type": "start_codon", "start": 6110, "end": 6112, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 6110, "end": 6148, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 6615, "end": 6683, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 6758, "end": 7040, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 7142, "end": 7319, "score": 1, "strand": 1, "phase": 2 }, { "type": "CDS", "start": 7411, "end": 7687, "score": 1, "strand": 1, "phase": 1 }, { "type": "CDS", "start": 7748, "end": 7850, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 7953, "end": 8098, "score": 1, "strand": 1, "phase": 2 }, { "type": "CDS", "start": 8166, "end": 8320, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 8419, "end": 8614, "score": 1, "strand": 1, "phase": 1 }, { "type": "CDS", "start": 8708, "end": 8811, "score": 1, "strand": 1, "phase": 0 }, { "type": "CDS", "start": 8927, "end": 9239, "score": 1, "strand": 1, "phase": 1 }, { "type": "CDS", "start": 9414, "end": 9494, "score": 1, "strand": 1, "phase": 0 }, { "type": "stop_codon", "start": 9492, "end": 9494, "strand": 1, "phase": 0 }, { "type": "three_prime_UTR", "start": 9495, "end": 9744, "score": 0.86, "strand": 1 } ] } ] }
Writing JBrowse Plugins
The JBrowse plugin system allows users to write their own JavaScript classes and AMD modules to extend JBrowse in nearly any way imaginable. Just a few things that can be done by plugins:
- Add new data store adapters to allow JBrowse to read data from existing web services.
- Add new track classes for custom track displays and behavior (WebApollo is one well-known plugin that does this).
- Add (or remove) options in the JBrowse dropdown menus.
- Add new types of track selectors.
Plugin Components
For an example plugin named MyPlugin, all of its files should be located in plugins/MyPlugin in the main JBrowse directory. There are no restrictions on what a plugin directory may contain.
A plugin is required to have a plugins/MyPlugin/js/main.js file, which is an AMD module that returns a JavaScript "class" that inherits from the JBrowse/Plugin class. JBrowse will create one instance of this class, which will persist during the entire time JBrowse is running. This class's constructor function is the entry point for your plugin's JavaScript code.
If a plugin contains custom CSS rules, it can optionally have a plugins/MyPlugin/css/main.css file as well, which JBrowse will load asynchronously. If your plugin code needs to know when the CSS is finished loading, the cssLoaded property of the plugin object contains a Dojo Deferred that is resolved when the CSS load is complete. Multiple CSS files can be loaded using @import statements in the main.css.
JBrowse also defines an AMD namespace that maps to your plugin's js directory, which your plugin code (and JBrowse itself) can use to load additional AMD modules and JavaScript classes. For example, our MyPlugin example could have a data-store adaptor in the file plugins/MyPlugin/js/Store/SeqFeature/FooBaseServices.js, which would be loaded with the module name "MyPlugin/Store/SeqFeature/FooBaseServices".
Example main.js
Here is an example plugin main.js, implemented using the Dojo declare class system.
define([ 'dojo/_base/declare', 'JBrowse/Plugin' ], function( declare, JBrowsePlugin ) { return declare( JBrowsePlugin, { constructor: function( args ) { var browser = this.browser; /* do anything you need to initialize your plugin here */ } }); });
Example plugin directory contents
plugins/MyPlugin/js plugins/MyPlugin/js/main.js plugins/MyPlugin/css plugins/MyPlugin/css/main.css plugins/MyPlugin/img plugins/MyPlugin/img/myimage.png