NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Difference between revisions of "JBrowse"
RobertBuels (Talk | contribs) (→Quick Start: Tutorial) |
RobertBuels (Talk | contribs) |
||
| Line 31: | Line 31: | ||
* libpng | * libpng | ||
| − | = | + | = Configuring JBrowse = |
| − | * [[JBrowse | + | * [[JBrowse Configuration Guide]] |
| − | ** [[JBrowse | + | ** [[JBrowse Configuration Guide#Reference Sequences|Formatting reference sequences]] (e.g. from FASTA files, or a Chado database) |
| − | ** [[JBrowse | + | ** [[JBrowse Configuration Guide#Feature Tracks|Creating feature tracks]] (e.g. from BED or GFF files, a Chado database, or the UCSC genome browser) |
| − | ** [[JBrowse | + | ** [[JBrowse Configuration Guide#Image Tracks|Creating image tracks]] (e.g. from WIG files) |
| − | ** [[JBrowse | + | ** [[JBrowse Configuration Guide#Searchable Names|Making features searchable by name]] |
| − | ** [[JBrowse | + | ** [[JBrowse Configuration Guide#Removing Tracks|Removing tracks]] |
* Additional topics: | * Additional topics: | ||
Revision as of 12:15, 11 April 2012
- Mature release
- Active development
- Active support
JBrowse is a genome browser with a fully dynamic AJAX interface. JBrowse renders most tracks using client-side JavaScript, with JSON as its data transfer format. JBrowse is being developed as the eventual official successor to GBrowse.
Contents
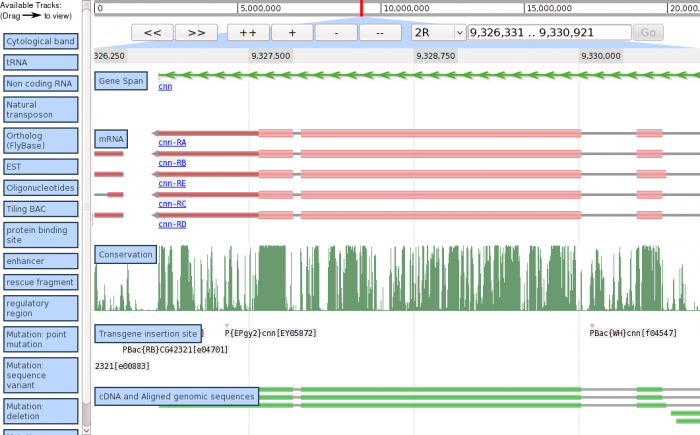
Demo
Quick Start: Tutorial
The Getting Started with JBrowse Tutorial provides a basic step-by-step recipe for quickly getting up and running with JBrowse.
Installation
See Installing JBrowse on Mac OS X and Linux for step-by-step installation instructions.
Prerequisites
- Perl modules from CPAN:
- BioPerl 1.6
- JSON
- JSON::XS (optional, for speed)
- Heap::Simple
- Heap::Simple::XS
- PerlIO::gzip
- Devel::Size
- libpng
Configuring JBrowse
- JBrowse Configuration Guide
- Formatting reference sequences (e.g. from FASTA files, or a Chado database)
- Creating feature tracks (e.g. from BED or GFF files, a Chado database, or the UCSC genome browser)
- Creating image tracks (e.g. from WIG files)
- Making features searchable by name
- Removing tracks
- Additional topics:
Contact
Please direct questions and inquiries regarding JBrowse to the mailing lists below.
In particular, requests for help should be directed to gmod-ajax@lists.sourceforge.net.
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| JBrowse | gmod-ajax | JBrowse help and general questions. | Nabble (2010/05+), Sourceforge |
| jbrowse-dev | JBrowse development discussions. | Nabble (2011/08+) |
JBrowse Development
JBrowse is an open-source project, started and managed by the laboratory of Ian Holmes at the University of California, Berkeley.
The JBrowse source code repository is kept on GitHub. Please feel very free to fork the code on GitHub and make modifications and improvements, submitting pull requests. GitHub has a very nice tutorial on how to get started with this style of development.
As of January 2012, the lead developer of JBrowse is Robert Buels. Most of JBrowse was originally written by Mitch Skinner.
Developer Communication
There is a mailing list for developers, and there is usually a teleconference on the 3rd Monday of the month at 2pm Pacific US time. We welcome participation from Please contact Robert Buels if you would like to listen in or participate.
Presentations
| Date | Presenter | Venue | Links |
|---|---|---|---|
| April 2012 | Robert Buels | GMOD 2012 Community Meeting | |
| January 2012 | Robert Buels | Plant and Animal Genome (PAG) XX | |
| April 2010 | Mitch Skinner | UCSC genome browser group ("genecats") meeting | |
| August 2009 | Ian Holmes | GMOD Community Meeting | Talk summary | PDF |
| January 2009 | Mitch Skinner | GMOD Community Meeting | Talk summary | PDF |
| July 2008 | Ian Holmes | GMOD Community Meeting | Talk summary | PDF |