NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Difference between revisions of "CMap"
(I get unreasonably irritated by irrelevant information being presented first) |
(→Screenshots) |
||
| Line 58: | Line 58: | ||
===Screenshots=== | ===Screenshots=== | ||
| − | Here are some sample maps from the [http://www.gramene.org/ Gramene Project]. | + | Here are some sample maps from the [http://www.gramene.org/ Gramene Project]. See also [http://www.gramene.org/cmap/entry.html other Gramene views]. |
| Line 77: | Line 77: | ||
[[Image:Cmap_sample3t.jpg]] | [[Image:Cmap_sample3t.jpg]] | ||
| − | |||
==Sites Using CMap== | ==Sites Using CMap== | ||
Revision as of 21:24, 15 December 2010
- Mature release
- Active development
- Active support
- CMap @ Gramene
- Tutorial @ Gramene
- Admin (perldoc)
- Other Doc
- Mailing List
- Download
- CMap3D
- 2008 Survey
Template:ComponentBoxEventsHeader
CMap is a web-based tool that allows users to view comparisons of genetic and physical maps. The package also includes tools for curating map data.
Contents
Requirements
- Perl 5.6.0 or higher
- A web server with CGI capabilites (e.g., Apache 1.x or 2.x)
- A relational database, e.g. MySQL (4.0+), Sybase Adaptive Server Enterprise, PostgreSQL, SQLite or Oracle (9.x)
- libgd 2.0.28 and GD.pm 2.15
- Various Perl modules available on CPAN
Documentation
- Gramene CMap User Tutorial,
- Available in both PDF and PowerPoint formats.
- CMap_API (SVN)
- README
- Changes
- INSTALL.pod
- ADMINISTRATION.pod
- CODE_OVERVIEW.pod
- TODO
- attributes-and-xrefs.pod
- CMAP Schema
- CMAP Tutorial at Texas A & M
- CMap Version 2 Design
Also see the CMap FAQ.
Presentations
- Genome Visualization and Comparison using CMap and Circos, poster from Genome Informatics 2009
- The CMap Comparative Map Browser and Displaying Population distributions with GBrowse and PhyloGeoViz from the GMOD Workshop at SMBE 2009
- CMap and CMAE from November 2007 GMOD Meeting
- CMap Version 0.16 from June 2006 GMOD Meeting
- CMap Update 2005 from May 2005 GMOD Meeting
- Gramene Comparative Genome Views: How Can Gramene Leverage Rice For The Other Grasses? poster form 2005 Plant and Animal Genomics meeting.
- CMap Update 2003 from September 2003 GMOD Meeting.
- CMap: Visualization of Comparative Maps from May 2003 GMOD Meeting.
Downloads
Demo & Screenshots
See the Gramene Project for a production version.
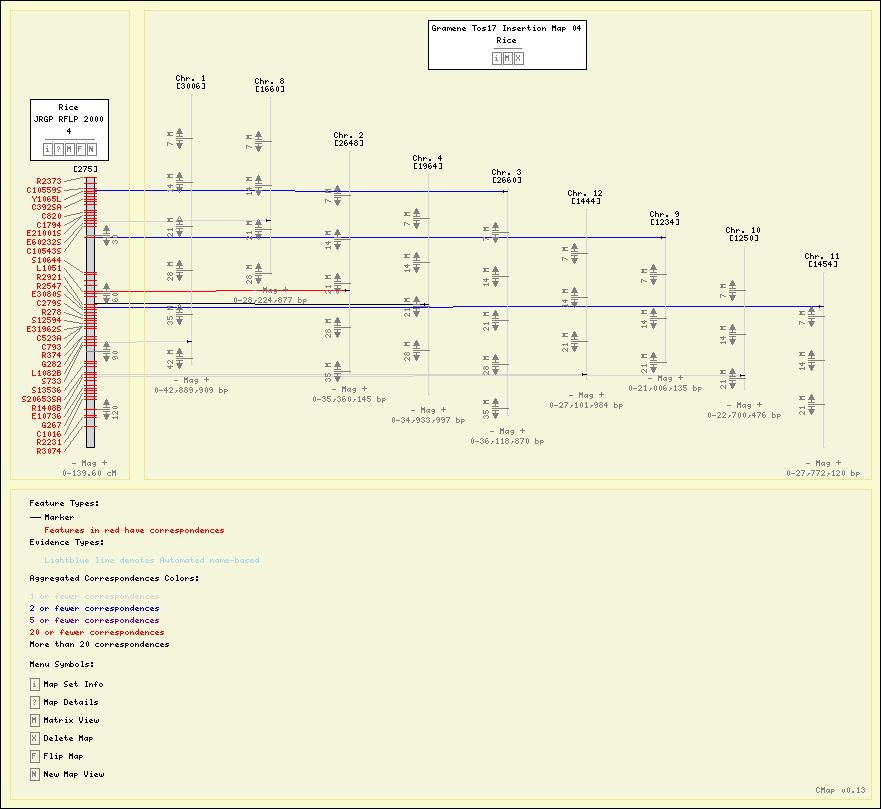
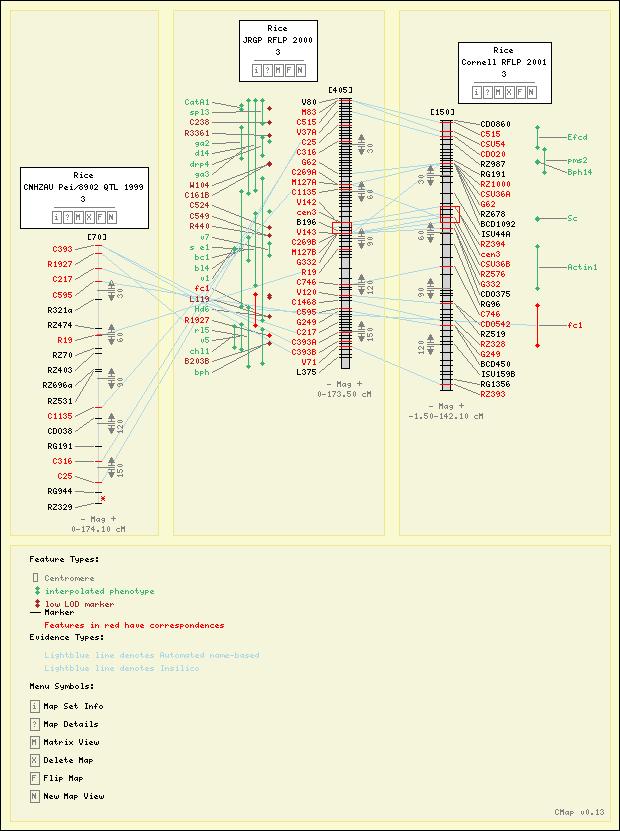
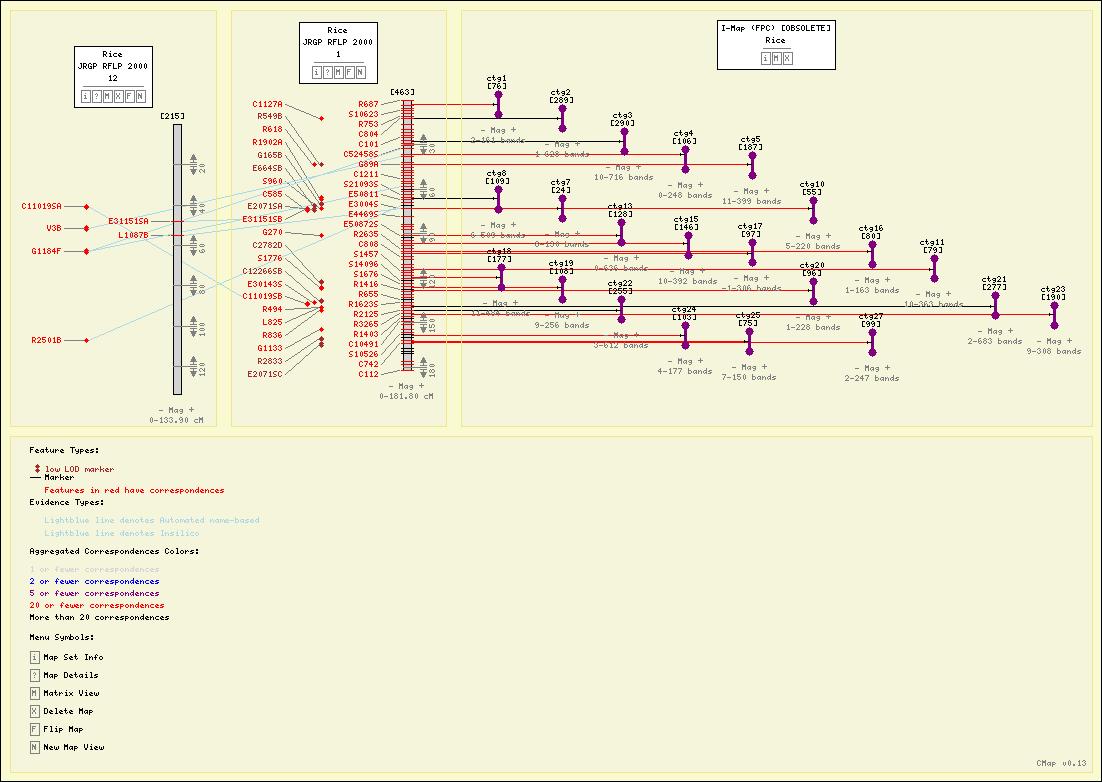
Screenshots
Here are some sample maps from the Gramene Project. See also other Gramene views.
Sites Using CMap
The following is a partial list of groups using CMap.
- Gramene: Comparative mapping resource for grasses (plant)
- NCGR's Legume Information System: A publicly accessible legume resource that will integrate genetic and molecular data from multiple legume species and enable cross-species comparisons
- BeeBase: The Honey Bee genome database
- BASC: Brassica database
- GrainGenes
- MAGI
- Genome Database for Rosaceae
- Cotton Marker Database
- SCRI Plant Bioinformatics Group
- Pristionchus pacificus Physical Map
- Comparitive Location Database
- Soybean Breeders Toolbox
- Composite Wheat Map
- PeanutMap
- CottonDB
- TropGene
- Populus at Oak Ridge
Code
Patches
- See the SourceForge Patch List.
- 1.0: Patched schema creation file for Postgres: {{SF_SVN|/cmap/trunk/sql/cmap.create.postgresql cmap.create.postgresql].
Mailing Lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| CMap | gmod-cmap | Discussion of CMap development, installation problems, etc. | Nabble (2010/05+), Sourceforge |
| gmod-cmap-commits | Notification of SVN activity for CMap. | Sourceforge |
Contact
Citation
CMap 1.01: a comparative mapping application for the Internet.
Ken Youens-Clark, Ben Faga, Immanuel V. Yap, Lincoln Stein and Doreen Ware, Bioinformatics 2009 25(22):3040-3042; doi:10.1093/bioinformatics/btp458.
Logo
The CMap logo was created by Kathy Bracken, a participant in the Spring 2010 Logo Program, while a design student at Linn-Benton Community College.
__NOTITLE__