GMOD
JBrowse/tool data
This page stores the data that populates the JBrowse wiki page.
| name = JBrowse | full_name = | contact_email = rbuels@gmail.com | status = mature | dev = active | support = active | type = Genome visualization | platform = web | logo = JBrowseLogo.png | input = GFF3, BED, FASTA, Wiggle, BigWig, BAM | output = | licence = LGPL and Artistic license | open source = yes | language = Javascript, Perl | release date = 2008 | home = http://jbrowse.org | demo_server = Volvox example data | wild_urls = | about = JBrowse is a genome browser with a fully dynamic HTML5 user interface, being developed as the successor to GBrowse. It is very fast and scales well to large datasets. JBrowse does almost all of its work directly in the user’s web browser, with minimal requirements for the server.
Features
- Fast, smooth scrolling and zooming. Explore your genome with unparalleled speed.
- Scales easily to multi-gigabase genomes.
- Supports GFF3, BED, FASTA, Wiggle, BigWig, BAM, and more.
- Very light server resource requirements. Serve huge datasets from a single low-cost cloud instance.
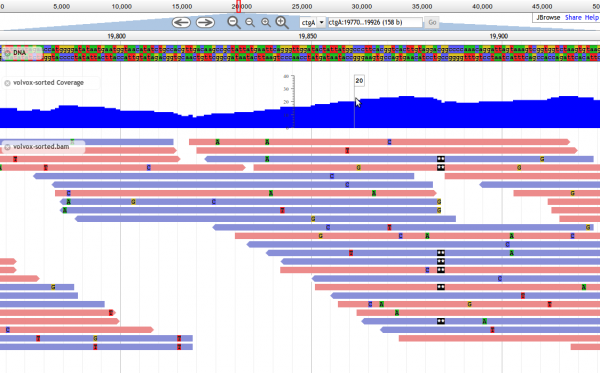
| screenshot =
| public_server = | dl = | dl_url = http://jbrowse.org/install/ | dl_src = | dl_src_url = http://github.com/GMOD/jbrowse | getting_started_preamble = The JBrowse Quick-Start Tutorial provides a basic step-by-step recipe for quickly getting up and running with JBrowse. | req = JBrowse requires libpng, Zlib, and GD development libraries, plus make and a C compiler. On Ubuntu, you can install these prerequisites using the command:
sudo apt-get install libpng-dev libgd2-noxpm-dev build-essential
For tips on installing these baseline libraries, see JBrowse Troubleshooting.
| install = The JBrowse Quick-Start Tutorial provides a basic step-by-step recipe for quickly getting up and running with JBrowse.
1. Download JBrowse onto your web server.
2. Unpack JBrowse into a directory that is served by your web browser.
On many systems, this defaults to /var/www.
cd /var/www
unzip JBrowse-*.zip
Make sure you have permissions to write to the contents of the
jbrowse/ directory you have just created.
3. Run the automated-setup script, ./setup.sh, which will attempt to
install all of JBrowse’s (modest) prerequisites for you in the
jbrowse/ directory itself. Note that setup.sh does not need to be
run as root or with sudo. For help troubleshooting failures of
setup.sh, see JBrowse
Troubleshooting.
4. Visit JBrowse on your machine, substituting the http://(your_machine/path_to_jbrowse)/index.html?data=sample_data/json/volvox. If you can see the included Volvox example data, you are ready to configure JBrowse to show your own data!
| config = See the JBrowse Configuration Guide for information on:
- Formatting reference sequences (e.g. from FASTA files, or a Chado database)
- Feature Tracks (e.g. from BED or GFF files, a Chado database, or the UCSC genome browser)
- Image Tracks (e.g. from WIG files)
- Wiggle/BigWig Tracks
- Name Search and Autocompletion
- Removing tracks
- Compressing data stored on the server
- URL control
- Faceted track selection
- Anonymous usage statistics
Additional topics: