GMOD
Galaxy Tutorial 2013
This walks you through setting up and running a Galaxy server. This tutorial will be taught by Dave Clements at the 2013 GMOD Summer School.
Contents
- 1 Open Ports 20-21, 8081
- 2 Starting AMI
- 3 Galaxy Project
- 4 Create a Galaxy instance
- 5 Running analyses with Galaxy
- 6 Administering Galaxy
- 7 Running Galaxy on the Cloud
Galaxy is a data integration and analysis framework for biomedical research. Galaxy allows nearly any tool that can be run from the command line to be integrated into it.
On top of these tools, Galaxy provides an accessible environment for interactive analysis that transparently tracks the details of analyses, a workflow system for convenient reuse, data management, sharing, publishing, and more.
Open Ports 20-21, 8081
For this tutorial, Galaxy will use port 8081. FTP, which is not used in this tutorial, but is used in Galaxy to upload large files, is on ports 20-21. Update your security group in AWS to open thee ports. Go to Security Groups, select the group you are using, and then click the Inbound tab and then add
| Create a new rule | Port range | Source |
|---|---|---|
| Custom TCP Rule | 8081 | 0.0.0.0/0 |
| Custom TCP Rule | 20-21 | 0.0.0.0/0 |
Click + Add rule after each, and then Apply Rule Changes after all have been added.
Starting AMI
Get the starting image for this tutorial - GMOD 2013 Galaxy Install Training Instance (ami-f1a1d998). It needs to be run as at least a large instance.
Galaxy Project
Some General Galaxy Resources
Before we get started, let’s highlight some Galaxy resources that may be useful to us along the way.
http://galaxyproject.org
The Galaxy Project home page
GalaxyWiki
All things Galaxy.
http://getgalaxy.org/
Hub page for installing and managing your own Galaxy instance.
http://usegalaxy.org/
The Galaxy project’s free public server.
Public Galaxy Servers
Current list of know publicly accessible Galaxy servers.
Galaxy Search
Integrated searches of all online Galaxy resources. Available searches:
Pan-Galactic Web Search
Search everything
Galaxy Mailing Lists Search
Search the (Nabble-powered) mailing list archives
Using Galaxy Search
Search online resources related to using Galaxy
Galaxy Admin and Development
Search
Search online resources related to deploying and developing Galaxy
Results from searches are often further broken down into categories
- All: give me everything
- Tools: show me doc on tools related to my search.
- Email: show email threads related to my search.
- Source code: show Galaxy source code related to my search
- Shared: Show published Galaxy objects related to my search
- Documentation: Show documentation (e.g. wiki pages, tool doc, …) related to my search.
- Abstracts: Show papers related to my search.
- Requests: Should feature requests related to my search.
This is all implemented using Google Custom Search.
Mailing Lists and Mailing Lists Search
Galaxy has several mailing lists, some of which are
very active
Learning hub page
Start here to learn how to use Galaxy.
Galaxy CiteULike group (@ CiteULike)
Seventeen different tags/categories on 1066+
publications
Create a Galaxy instance
See http://getgalaxy.org.
Prerequisites
You need a Python interpreter, version 2.6 or greater. Python 3 is a different language and is currently not supported. The Amazon Machine Image (AMI) used for this course includes version 2.7.3 of the interpreter.
$ python --version
Python 2.7.3
Galaxy is distributed (and developed) using a distributed version control system called Mercurial. The AMI already includes mercurial version 2:0.2:
$ hg --version
Mercurial Distributed SCM (version 2.0.2)
...
Galaxy needs a newer version of bedtools
bedtools is installed on this machine, but it’s an
older version without the
bedtools executable in it, and Galaxy needs that. Remove the existing
bedtools package, and install
a newer one.
DO NOT DO THIS NOW as it has already been done on your image:
Remove the old version, get the new version, unpack and compile it, and then put it where it can be found.
$ sudo apt-get remove bedtools
$ cd ~/Galaxy
$ curl http://bedtools.googlecode.com/files/BEDTools.v2.17.0.tar.gz > BEDTools.tar.gz
$ tar -zxf BEDTools.tar.gz
$ cd bedtools-2.17.0/
$ make
$ sudo cp -i bin/* /usr/bin
Clone the Galaxy repository
The development and release repositories are available through the bitbucket hosting service.
DO NOT DO THIS NOW as it has already been done on your image:
To create a local clone of the release repository run the following:
$ cd ~/Galaxy
$ hg clone http://bitbucket.org/galaxy/galaxy-dist
$ cd galaxy-dist
$ hg update stable
Update Galaxy Configuration File
Often you can just fire up Galaxy at this point. However, we want a few
things to be different from the default installation. Galaxy’s main
configuration file is universe_wsgi.ini. By default, that file is
created at initialization time by copying universe_wsgi.ini.sample.
However, if the file already exists it is not copied over. Copy the file
and update it:
$ cd ~/Galaxy/galaxy-dist
$ cp universe_wsgi.ini.sample universe_wsgi.ini
$ nano universe_wsgi.ini
Change the port from
#port = 8080
to this:
port = 8081
Galaxy, like WebApollo and several other components that are also covered at the course, will listen to port 8080 by default; to avoid stomping on that earlier work, we will configure Galaxy to listen to a different port.
Change the host from
#host = 127.0.0.1
to:
host = 0.0.0.0
This makes Galaxy visible to remote hosts, such as your laptop
Set the brand to make it obvious that you are working on your Galaxy instance
Change this:
#brand = None
to this:
brand = My Super Cool Brand
Actually use something shorter.
Use a more robust database
Out of the box Galaxy includes the embedded SQLite database. This allows Galaxy to run with zero-configuration and provides an excellent solution for single-user Galaxy installations being used for tool development. However, for any multi-user scenario a more robust database will be needed for Galaxy to be reliable. We highly recommend Postgres, although other databases are known to work. Postgres is already installed on our AMI (it’s the default DBMS for Chado)
Update universe_wsgi.ini file to use Postgres. Update the
database_connection in your Galaxy config file to look like:
# -- Database
# By default, Galaxy uses a SQLite database at 'database/universe.sqlite'. You
# may use a SQLAlchemy connection string to specify an external database
# instead. This string takes many options which are explained in detail in the
# config file documentation.
#database_connection = sqlite:///./database/universe.sqlite?isolation_level=IMMEDIATE
database_connection = postgres://ubuntu:@localhost:5432/galaxydb
Save the file.
The ubuntu user has permission to create databases, so let’s create
the database that we told Galaxy to connect to:
$ createdb galaxydb
Run, Galaxy, Run!
Galaxy includes a script to run it. This script also performs the Galaxy initialization the first time it is run. Run it now:
$ sh run.sh
Initializing external_service_types_conf.xml from external_service_types_conf.xml.sample
Initializing migrated_tools_conf.xml from migrated_tools_conf.xml.sample
Initializing reports_wsgi.ini from reports_wsgi.ini.sample
Initializing shed_tool_conf.xml from shed_tool_conf.xml.sample
Initializing tool_conf.xml from tool_conf.xml.sample
... (several minutes pass while install rolls through database changes) ...
galaxy.webapps.galaxy.buildapp DEBUG 2013-07-15 18:52:06,052 Enabling 'x-forwarded-host' middleware
galaxy.webapps.galaxy.buildapp DEBUG 2013-07-15 18:52:06,053 Enabling 'Request ID' middleware
Starting server in PID 7158.
serving on 0.0.0.0:8081 view at http://127.0.0.1:8081
This script performs several significant actions the first time it is run:
- Creates initial configuration files and empty directories for storing data files
- Fetches all of the Galaxy framework’s dependencies, packaged as Python eggs, for the current platform.
- Initializes its database. Galaxy uses a database migration system to automatically handle any changes to the database schema. On first load it runs all migrations to ensure the database is in a known state, which may take a little time.
Once the database is initialized, the normal startup process proceeds, loading tool configurations, starting the job runner, and finally initializing the web interface on the requested port. You can now access your Galaxy at http://ec2-##-##-##-##.compute-1.amazonaws.com:8081.
Running analyses with Galaxy
See also Galaxy 101 tutorial
Without any additional configuration, there is already a lot we can do with our first Galaxy instance. As an example, let’s work through an analysis that is based on, but distinct from the Galaxy 101 tutorial.
1. Access your new Galaxy instance
Start a web browser and access http://ec2-##-##-##-##.compute-1.amazonaws.com:8081.
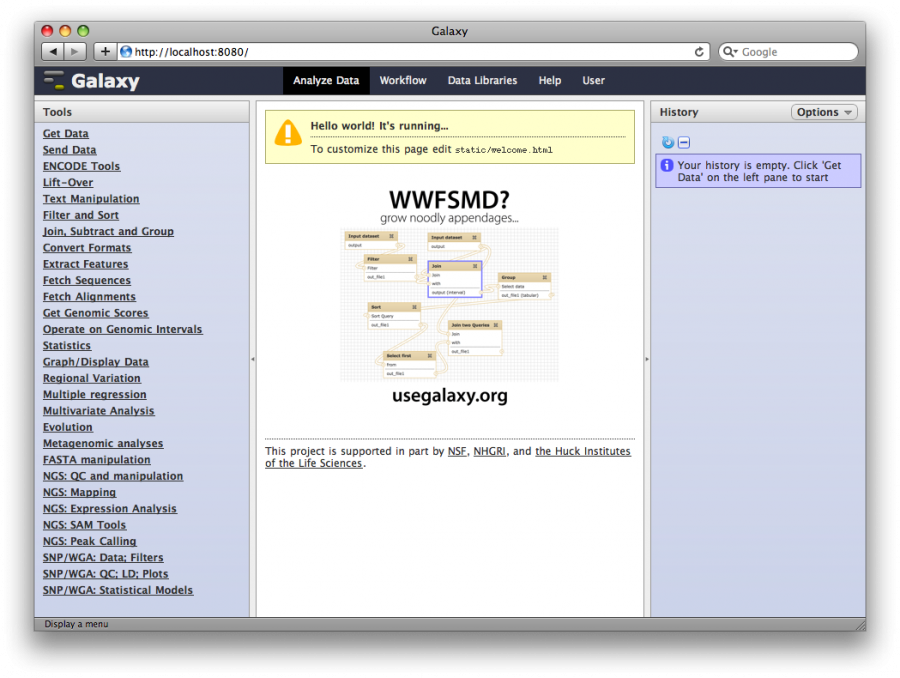
2. Create a user
In the top bar, select User → Register. Enter your
- Email address
- Password (use a low-security password, it’s going over the net unencrypted)
- Public name: Public names must be at least four characters in length and contain only lower-case letters, numbers, and the ‘-‘ character.
and click Submit.
Registering is not required in order to use Galaxy. However, to use all of it, users need to register.
3. Lets answer a question
Now that Galaxy is up and running, let’s use it to answer a question
We scooped up an unknown beast out of the slime and sent it off to the sequencing core to get it sequenced and an assembly so we can study it. Turns out it’s an archaea, and it’s new. We have subsequently run the assembly through a gene prediction pipeline, and have also identified potential transcription binding sites (TFBSs) using another pipeline.
What we now want to know is: which exons/genes have the most overlapping / embedded TFBSs?
4. Get the data into Galaxy
Select Tools → Get Data → Upload Data. This brings up the upload data form from which you can
- upload data from your computer, or
- upload data from one or more URLs
- cut and paste data directly
- bring files into your workspace that you have previously send to Galaxy via FTP.
We are going to use the URL option. Cut and paste these URLs into the URL/Text box
http://bx.psu.edu/~clements/Events/GMOD2013/m.vannielli.TFBs.bed
http://bx.psu.edu/~clements/Events/GMOD2013/m.vannielli.sequence.fasta
http://bx.psu.edu/~clements/Events/GMOD2013/m.vannielli.wholegene.bed
and click Execute.
This will import those 4 datasets your history. Let’s take a look at the data. For each dataset,
- Click on the dataset name for a preview.
- Poke the eye to see the full dataset.
- Click on pencil icon and give each dataset a better name (like
M vannielli Exons). Click Save. - Change the history name from
unnamed history(which is true, but not useful) to something more meaningful.
5. Identify genes and TFBSs that overlap
Select Tools → Operate on Genomic Intervals → Join.
Join genes with TFBS, with min overlap of 1 bp. Return Only records that are joined (INNER JOIN).
Takes the two bed files and joins them together into double-wide records where the first n columns are from the genes dataset and the last columns are from the repeats dataset. Furthermore, it only creates records when an gene and a TFBS overlap.
Take a close look at the dataset. Note that
- Some genes were dropped
- Some TFBS were dropped
- Some genes occur multiple times
Make sure you understand why.
6. Group and Count
Now we want to walk through the pairings and count the number of times each gene occurs. This number is the number of TFBS that overlap with each gene.
We are going to do another operation that is borrowed from relational databases. Select Tools → Join, Subtract, and Group → Group.
Select the gene-TFBS pairings dataset and set Group by column to c4, the column in the dataset that contains the gene name.
Then click Add new operation and then set Type to Count.
This tells Galaxy to walk through the dataset, create a group for each different value of column 4 (the gene name), and then count the number of records that were in that group (i.e. the number of records that had each gene name).
This produces a two column dataset. The first column is the value of the column we grouped by. The second is the number of records in the dataset that have that gene name.
If we were to now to run Tools → Filter and Sort → Sort on this dataset, we would have the answer to our original question:
Which genes have the most overlapping TFBS?
We have the list of genes, and the counts in them. We could use this dataset in further analysis, email it someone, etc..
7. Get Gene Info back
However, we can do better. We have lost some information about the genes (like position, strand, and so on) that we had in the original gene dataset. If we can reclaim that information, and add to it, we can produce a more useful dataset that we can visualize right now.
The original gene dataset downloaded from UCSC had a meaningless score column. Let’s replace that with the repeat count.
First, bring the original gene information together with the counts.
Select Tools → Join, Subtract and Group → Join two Datasets. Set the first dataset to Genes with repeat counts and the second to be the original Genes dataset.
Join them using column c1 and column c4, which are the gene names in both datasets.
This produces and a dataset with the gene repeat counts in the first two columns and the gene information in the rightmost columns.
Now, use the Cut tool to reshuffle these columns into a valid 6 column BED file with the repeat count in column 5, the score column.
Select Tools → Text Manipulation → Cut. Enter
c3,c4,c5,c6,c2,c8
Finally, note that even though we now have what is a valid 6 column bed file, Galaxy doesn’t know that. We need to tell it. We should also give the dataset a better name. Click on the datasets’ pencil icon and set the dataset name to something meaningful. Then click on the Datatype type and select bed from the pulldown.
8. Visualize Your Analysis
We can now visualize our results in Galaxy. Click on the visualize icon and select Trackster’ from the pulldown, and then click on View in new visualization.
Define this assembly to Galaxy
You may have noticed that In the preview of each dataset it shows
database: ?
The database (also referred to as a build) specifies which genome assembly this dataset is associated with. A genome assembly is usually named with an abbreviation for the species and a version number. For example, “mm9” represents the Mus musculus 9” assembly released in 2007, and “mm10” is the more recent assembly released in 2010. If these datasets had been directly imported into Galaxy from, for example, the UCSC Table Browser, or BioMart then the database for these datasets would have been set automatically.
The “?” means that Galaxy does not know which genome assembly the datasets (or more precisely, the coordinates in the datasets) are associated with. Genome assemblies are defined to Galaxy by the instance’s administrators. Most Galaxy servers know about widely used assemblies, and if our datasets were for mm9 or mm10 (or any of many other choices), we (as users) could just tell Galaxy that and be done.
However, these datasets aren’t from a common assembly - they are from your sequencing center and are for some novel archaea that was pulled out of a mudflat (that is a bald-faced lie; it is actually Methanococcus vannielii and this data came from the UCSC table browser, but we will pretend not to know that).
One feature of Galaxy is that users can define their own custom builds/genomes to the system. Once defined, they will be there every time that user logs in in the future.
When you create a visualization, you are asked if you want to use a known build, or if you want to create a custom build. Since we have a custom build, select that option. Under New Build define
- Name (eg: Hamster):
- Key (eg: hamster_v1):
and select the FASTA file from your history. Click Submit. Now, every time you log into this server in the future, Galaxy will know about this assembly. No click on the final datasets visualization icon again.
And …
Our hopes are crushed:
Trackster Error
/bin/sh: 1: bedGraphToBigWig: not found
Do a web search for bedGraphToBigWig. Eventually that will lead you to UCSC, where you can download it and install it.
cd ~/Galaxy
curl http://hgdownload.cse.ucsc.edu/admin/exe/linux.x86_64/bedGraphToBigWig > bedGraphToBigWig
chmod 555 bedGraphToBigWig
sudo cp bedGraphToBigWig /usr/bin/
Click Try again. Voillà.
Administering Galaxy
1. Make yourself an admin
On the command line, stop Galaxy by entering <ctrl-c>. Then edit
universe_wsgi.ini file and add yourself as an administrator:
# users (email addresses). These users will have access to the Admin section
# of the server, and will have access to create users, groups, roles,
# libraries, and more. For more information, see:
# http://wiki.g2.bx.psu.edu/Admin/Interface
#admin_users = None
admin_users = your@email.adr
2. Enable Tool Shed
While we are in universe_wsgi.ini, lets also enable this server to use
the Galaxy Tool Shed.
Define:
# Tool config files, defines what tools are available in Galaxy.
# Tools can be locally developed or installed from Galaxy tool sheds.
#tool_config_file = tool_conf.xml,shed_tool_conf.xml
tool_config_file = tool_conf.xml,shed_tool_conf.xml
# Default path to the directory containing the tools defined in tool_conf.xml.
# Other tool config files must include the tool_path as an attribute in the <toolbox> tag.
#tool_path = tools
tool_path = tools
Save the changes and then in the shell enter
$ mkdir ../shed_tools
This is the default location for tools installed through the tool shed.
Now, restart Galaxy:
$ sh run.sh
3. Find and install a Venn Diagram drawing tool
Once your Galaxy has restarted, reload your web page. You should now see an Admin link in the top menu bar. Click on it, and then Tool sheds → Search and browse tool sheds → Galaxy Main Tool Shed. Search for “venn”.
An error occurred with this dataset: Traceback (most recent call last): File “/home/ubuntu/Galaxy/shed_tools/toolshed.g2.bx.psu.edu/repos/peterjc/venn_list/c96bef0643dc/venn_list/tools/plotting/venn_list.py”, line 13, in import rpy ImportError: No module named rpy
Do a web search for rpy debian package
apt-get -s install python-rpy
sudo apt-get install python-rpy
An error occurred with this dataset: Requires the R library limma (for vennDiagram function)
Do a web search for limma debian package. Pick the one that involves R and biology.
sudo apt-get install r-bioc-limma
Success.
Running Galaxy on the Cloud
Go to CloudLaunch