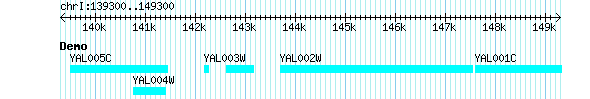NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Glyphs and Glyph Options
Contents
- 1 Selecting and Configuring Glyphs
- 2 Values Used in Glyph Options
- 3 Glyphs and their Configuration Options
- 3.1 generic
- 3.2 alignment
- 3.3 allele_tower
- 3.4 anchored_arrow
- 3.5 arrow
- 3.6 box
- 3.7 broken_line
- 3.8 cds
- 3.9 christmas_arrow
- 3.10 cross
- 3.11 dashed_line
- 3.12 diamond
- 3.13 dna
- 3.14 dot
- 3.15 dumbbell
- 3.16 ellipse
- 3.17 ex
- 3.18 extending_arrow
- 3.19 fixedwidth
- 3.20 flag
- 3.21 gene
- 3.22 graded_segments
- 3.23 group
- 3.24 hat
- 3.25 heat_map
- 3.26 heat_map_ideogram
- 3.27 heterogeneous_segments
- 3.28 hidden
- 3.29 hybrid_plot
- 3.30 ideogram
- 3.31 image
- 3.32 lightning
- 3.33 line
- 3.34 merge_parts
- 3.35 merged_alignment
- 3.36 minmax
- 3.37 oval
- 3.38 pairplot
- 3.39 pentagram
- 3.40 phylo_align
- 3.41 pinsertion
- 3.42 primers
- 3.43 processed_transcript
- 3.44 protein
- 3.45 ragged_ends
- 3.46 rainbow_gene
- 3.47 redgreen_box
- 3.48 redgreen_segment
- 3.49 repeating_shape
- 3.50 rndrect
- 3.51 ruler_arrow
- 3.52 saw_teeth
- 3.53 segmented_keyglyph
- 3.54 segments
- 3.55 smoothing
- 3.56 span
- 3.57 spectrogram
- 3.58 splice_site
- 3.59 stackedplot
- 3.60 ternary_plot
- 3.61 text_in_box
- 3.62 three_letters
- 3.63 tic_tac_toe
- 3.64 toomany
- 3.65 trace
- 3.66 track
- 3.67 transcript
- 3.68 transcript2
- 3.69 translation
- 3.70 triangle
- 3.71 two_bolts
- 3.72 vista_plot
- 3.73 wave
- 3.74 weighted_arrow
- 3.75 whiskerplot
- 3.76 wiggle_box
- 3.77 wiggle_density
- 3.78 wiggle_xyplot
- 3.79 wiggle_minmax
- 3.80 xyplot
See also: GBrowse Configuration/Glyphs
GBrowse offers many different options for displaying tracks. Tracks are generated by the Perl Bio::Graphics module, which is the definitive reference for the various display options in GBrowse. This page provides an introduction to the most commonly used glyphs.This page or section needs to be edited. Please help by editing this page to add your revisions or additions.
Selecting and Configuring Glyphs
Every GBrowse [Track] stanza should have a glyph option. This option selects the shape and behavior of each feature rendered in the track. Other options fine tune the glyph's appearance by selecting its height, color, and other display properties. Several standard display options are shared among all glyphs; others are glyph-specific.
The simplest [Track] stanza will look like this:[Demo] feature = gene glyph = generic
The "generic" glyph is the basis of all other glyphs and simply draws a solid box using the cyan color. A track rendered with this configuration will look like Figure 1.
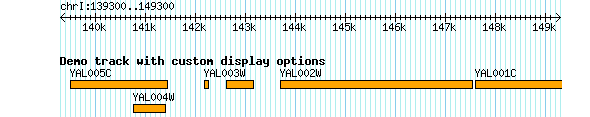
You can change the default appearance of the generic glyph using the standard height, bgcolor, and fgcolor options. You can also assign a longer key to the track using the key option. For example:[Demo] feature = gene glyph = generic bgcolor = orange fgcolor = black height = 9 key = Demo track with custom display options
Figure 2 shows how this will appear.
Values Used in Glyph Options
Before we dive into the various glyphs and their options, we will discuss colors, true/false flags, and other values that are used in the options.
Colors
Many options accept color values. You can specify colors by name, by HTML value, or using CSS (cascading stylesheet) notation.
By Name
GBrowse color-related options accept human readable color names like "yellow" and "chartreuse". The full list is contained in the color chart in Figure 3.By HTML Value
GBrowse accepts HTML style color values in the form "#RRGGBB", where RR, GG and BB are the red, green and blue intensity values respectively. The intensity values are expressed in hexadecimal, and range from 00 (completely off) to FF (completely on). Black is #000000 (all colors off), and white is #FFFFFF (all colors at full intensity). See HTML Colors for several nice tables of common #RRGGBB combinations. For example, to give a glyph a bright blue background, you could specify a bgcolor option like this:
bgcolor = #0000FF
Using CSS Notation
GBrowse also accepts Cascading Stylesheeet Level 3 color specifications, which are in the form "rgb(r,g,b)", where r, g, b are red, green and blue color intensities expressed as percentages or decimal values from 0 to 255. In the percentage form, a bright red background color would be expressed as:
bgcolor = rgb(100%,0%,0%)
in the second form, the color yellow (which is an equal combination of red and green in rgb space) could be written as:
bgcolor = rgb(255,255,0)
Alpha (Transparency) Values
You can also create colors with various degrees of translucency by specifying an alpha value. Alpha values can be specified as a hexadecimal value, as a floating point number between 0.0 and 1.0, or as a percentage from 0% to 100%. When using a named color, you can specify the degree of opacity in either of these ways:
bgcolor = blue:0.8 bgcolor = blue:80%
Either of these forms specifies a blue value that is 80% opaque. Values of 0.0 or 0% will create an entirely transparent color (which will appear as a "hole" in the drawing canvas), while values of 1.0 or 100% will create an entirely opaque color.
To specify alpha values with CSS syntax, use the "rgba(r,g,b,a)" notation, where "a" is the value of the alpha channel expressed either as a floating point number between 0.0 and 1.0, or as a percentage. In this form, 80% blue can be expressed in any of these forms:
bgcolor = rgba(0%,0%,100%,80%) bgcolor = rgba(0,0,255,80%) bgcolor = rgba(0,0,255,0.8)
Finally, you can use an extension of the HTML color format to specify the color and alpha values as #RRGGBBAA, where AA is the hexadecimal representation of alpha. This format is a bit unintuitive because the alpha channel is expressed as a hexadecimal number between 0 and 7F (decimal 127), where 0 is completely opaque and 7F is completely transparent. This reversal of the direction of opacity reflects how alpha is represented in the underlying graphics data structure. In this format, 80% blue can be expressed as:
bgcolor = #0000FF19
True/False Values
Many options are flags that turn certain features on or off. GBrowse follows the Perl programming language convention of using 0 (zero) as false, and accepting any non-zero value as true. Most commonly, people use 0 for false and 1 for true. For example, the "gene" glyph takes a decorate_introns option, which if true draws little arrowheads on the introns of the gene's transcripts to indicate the direction of transcription. To activate this feature, set the option to a true value:
decorate_introns = 1
to deactivate, set it to false:
decorate_introns = 0
Fonts
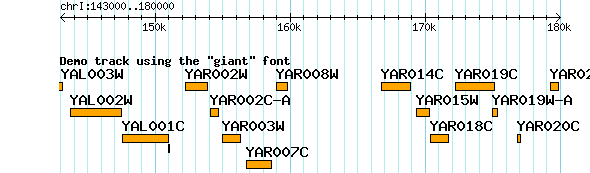
Some options, such as the generic font option, expect a font name. GBrowse provides a small set of fixed-width fonts provided by the [GD library package], named "gdTinyFont","gdSmallFont", "gdMediumBoldFont","gdLargeFont" and "gdGiantFont". They are all shown together in Figure 4.If not specified, most glyphs default to "gdSmallFont". Figure 5 shows the effect of changing the font to "gdLargeFont" and turning on feature labeling, as shown in the example below:
[Demo] feature = gene glyph = generic font = gdGiantFont bgcolor = orange fgcolor = black height = 9 key = Demo track using the "giant" font
Screen Measurements
All dimensions, such as height and margins, are given in pixel coordinates:
height = 12
No other measurement types, such as points, centimeters or ems are recognized.
Glyphs and their Configuration Options
Ths section describes each of the common glyphs, shows a sample configuration stanza, and lists its options and possible values. A list of some of the defined glyphs can be found in the documentation of Bio::Graphics.
You can find a sizable list of samples of each glyph at webgbrowse.cgb.indiana.edu.
generic
The generic glyph is the basis for all other glyphs and is the default if the glyph option is not explicitly stated. The following is a representative track stanza that uses several of generic's possible options:
[Demo]
feature = gene
glyph = generic
font = gdSmallFont
bgcolor = orange
fgcolor = black
linewidth = 2
height = 9
stranded = 1
label = 1
connector = solid
description = sub {my $feature = shift;
my @gene_names = $feature->get_tag_values('gene');
return "Gene @gene_names";
}
link = http://www.google.com/search?q=$name
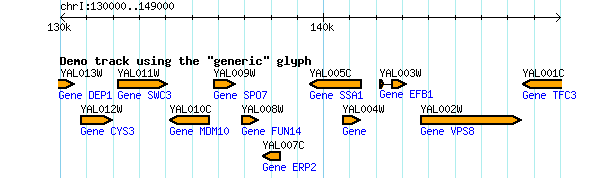
key = Demo track using the "generic" glyph
The result is shown in Figure 6. This example illustrates a pair of advanced features described in the GBrowse 2.0 HOWTO. The first advanced feature is the link option, which specifies where to link to when the user clicks on the feature. The example specifies "http://www.google.com/search?q=$name". When GBrowse renders the page, $name will be replaced with the name of the feature as specified in the underlying database.
A more complex example is the description option, which controls the optional text printed below the feature, is a Perl subroutine callback. It retrieves the feature from the subroutine parameter list (using the "shift" operator) and stores it into a variable named "$feature". It then calls the feature's get_tag_values() method to retrieve descriptive tags of type "gene", here used to store the common gene name. The gene names are stored into an array named "@gene_names." Lastly, the subroutine returns a string consisting of the word "Gene" followed by the list of gene names.
The full list of options recognized by generic and all descendent glyphs is given in the following table:
| Name | Description | Default Value |
|---|---|---|
| key | Description of the track in the tracks table. | same as the stanza label |
| category | The category of the track in the tracks table. You may use the format "category1:category2:category3" to specify subcategories. | Uncategorized tracks will be placed in a hold-all named "General" |
| fgcolor | The foreground color of the glyph; this will be used in drawing the rectangular border of the glyph. | black |
| bgcolor | The background color of the glyph; this will be used in filling the contents of the glyph rectangle. | cyan |
alignment
Bio::Graphics::Glyph::alignment
allele_tower
Bio::Graphics::Glyph::allele_tower
anchored_arrow
Bio::Graphics::Glyph::anchored_arrow
arrow
Bio::Graphics::Glyph::arrow
box
Bio::Graphics::Glyph::box
broken_line
Bio::Graphics::Glyph::broken_line
cds
Bio::Graphics::Glyph::cds
christmas_arrow
Bio::Graphics::Glyph::christmas_arrow
cross
Bio::Graphics::Glyph::cross
dashed_line
Bio::Graphics::Glyph::dashed_line
diamond
Bio::Graphics::Glyph::diamond
dna
Bio::Graphics::Glyph::dna
dot
Bio::Graphics::Glyph::dot
dumbbell
Bio::Graphics::Glyph::dumbbell
ellipse
Bio::Graphics::Glyph::ellipse
ex
Bio::Graphics::Glyph::ex
extending_arrow
Bio::Graphics::Glyph::extending_arrow
fixedwidth
Bio::Graphics::Glyph::fixedwidth
flag
Bio::Graphics::Glyph::flag
gene
Bio::Graphics::Glyph::gene
graded_segments
Bio::Graphics::Glyph::graded_segments
group
Bio::Graphics::Glyph::group
hat
Bio::Graphics::Glyph::hat
heat_map
Bio::Graphics::Glyph::heat_map
heat_map_ideogram
Bio::Graphics::Glyph::heat_map_ideogram
heterogeneous_segments
Bio::Graphics::Glyph::heterogeneous_segments
Bio::Graphics::Glyph::hidden
hybrid_plot
Bio::Graphics::Glyph::hybrid_plot
ideogram
Bio::Graphics::Glyph::ideogram
image
Bio::Graphics::Glyph::image
lightning
Bio::Graphics::Glyph::lightning
line
Bio::Graphics::Glyph::line
merge_parts
Bio::Graphics::Glyph::merge_parts
merged_alignment
Bio::Graphics::Glyph::merged_alignment
minmax
Bio::Graphics::Glyph::minmax
oval
Bio::Graphics::Glyph::oval
pairplot
Bio::Graphics::Glyph::pairplot
pentagram
Bio::Graphics::Glyph::pentagram
phylo_align
Bio::Graphics::Glyph::phylo_align
pinsertion
Bio::Graphics::Glyph::pinsertion
primers
Bio::Graphics::Glyph::primers
processed_transcript
Bio::Graphics::Glyph::processed_transcript
protein
Bio::Graphics::Glyph::protein
ragged_ends
Bio::Graphics::Glyph::ragged_ends
rainbow_gene
Bio::Graphics::Glyph::rainbow_gene
redgreen_box
Bio::Graphics::Glyph::redgreen_box
redgreen_segment
Bio::Graphics::Glyph::redgreen_segment
repeating_shape
Bio::Graphics::Glyph::repeating_shape
rndrect
Bio::Graphics::Glyph::rndrect
ruler_arrow
Bio::Graphics::Glyph::ruler_arrow
saw_teeth
Bio::Graphics::Glyph::saw_teeth
segmented_keyglyph
Bio::Graphics::Glyph::segmented_keyglyph
segments
Bio::Graphics::Glyph::segments
smoothing
Bio::Graphics::Glyph::smoothing
span
Bio::Graphics::Glyph::span
spectrogram
Bio::Graphics::Glyph::spectrogram
splice_site
Bio::Graphics::Glyph::splice_site
stackedplot
Bio::Graphics::Glyph::stackedplot
ternary_plot
Bio::Graphics::Glyph::ternary_plot
text_in_box
Bio::Graphics::Glyph::text_in_box
three_letters
Bio::Graphics::Glyph::three_letters
tic_tac_toe
Bio::Graphics::Glyph::tic_tac_toe
toomany
Bio::Graphics::Glyph::toomany
trace
Bio::Graphics::Glyph::trace
track
Bio::Graphics::Glyph::track
transcript
Bio::Graphics::Glyph::transcript
transcript2
Bio::Graphics::Glyph::transcript2
translation
Bio::Graphics::Glyph::translation
triangle
Bio::Graphics::Glyph::triangle
two_bolts
Bio::Graphics::Glyph::two_bolts
vista_plot
This is a combination XY or density plot with superimposed peak calls; it is typically used with ChIP-seq and ChIP-chip data. Please see Using the vista_plot Glyph for instructions.
wave
Bio::Graphics::Glyph::wave
weighted_arrow
Bio::Graphics::Glyph::weighted_arrow
whiskerplot
Bio::Graphics::Glyph::whiskerplot
wiggle_box
Bio::Graphics::Glyph::wiggle_box
wiggle_density
A density glyph compatible with the "wig" data format. Varies the intensity of the hues corresponding to the value of the data. You can set a pivot point around which the color changes to more easily visualize negative values.
[wiggle_density] feature = Mooney_wiggle glyph = wiggle_density height = 10 fgcolor = black bicolor_pivot = 0 pos_color = red neg_color = blue key = Mooney ChIP-chip (Density) category = Experimental Data label = 1
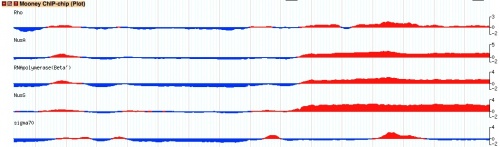
wiggle_xyplot
An xy-plot compatible with dense "wig" data. See the documentation for Bio::Graphics::Glyph::wiggle_xyplot.
It produces the typical xyplot, but uses the wig data for dense data sets. In the example the bicolor_pivot parameter has been set to 0, and the pos_color and neg_color were set to obtain the conditional coloring.
[wiggle_xyplot]
feature = Mooney_wiggle
glyph = wiggle_xyplot
height = 20
fgcolor = black
bicolor_pivot = 0
pos_color = red
neg_color = blue
key = Mooney ChIP-chip (Plot)
category = Experimental Data
label = 0
citation = Mooney RA, Davis SE, Peters JM, Rowland JL et al.
Regulator trafficking on bacterial transcription units in vivo. Mol Cell 2009 Jan 16;33(1):97-108.
label = 1
wiggle_minmax
Bio::Graphics::Glyph::wiggle_minmax
xyplot
Bio::Graphics::Glyph::xyplot