NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
Difference between revisions of "Chado Natural Diversity Module"
(→Table: nd_experiment_contact) |
(→Table: nd_experiment_contact) |
||
| Line 93: | Line 93: | ||
==Table: nd_experiment_contact== | ==Table: nd_experiment_contact== | ||
| − | primary contact / submitter of these nd_experiments (nd, where assays are not submitted separately this may be better stored in project_contact) | + | primary contact / submitter of these nd_experiments (nd, where assays are not submitted separately this may be better stored in project_contact). |
{| border="1" cellpadding="3" | {| border="1" cellpadding="3" | ||
Revision as of 17:21, 16 December 2010
The Chado Natural Diversity Module is an extension to the Chado schema to better support natural diversity data.
Eventually this page will resemble the other Chado Module pages, with an overview followed by a detailed explanation of the tables, columns, and relationships in the module. However, while the module is under development, this page will have an alternative structure.
Contents
- 1 Introduction
- 2 Use Cases
- 3 ER Diagram
- 4 Tables
- 4.1 Table: nd_experiment
- 4.2 Table: nd_experiment_contact
- 4.3 Table: nd_experiment_dbxref
- 4.4 Table: nd_experiment_genotype
- 4.5 Table: nd_experiment_phenotype
- 4.6 Table: nd_experiment_project
- 4.7 Table: nd_experiment_protocol
- 4.8 Table: nd_experiment_pub
- 4.9 Table: nd_experiment_stock
- 4.10 Table: nd_experiment_stock_dbxref
- 4.11 Table: nd_experiment_stockprop
- 4.12 Table: nd_experimentprop
- 4.13 Table: nd_geolocation
- 4.14 Table: nd_geolocationprop
- 4.15 Table: nd_protocol
- 4.16 Table: nd_protocol_reagent
- 4.17 Table: nd_protocolprop
- 4.18 Table: nd_reagent
- 4.19 Table: nd_reagent_relationship
- 4.20 Table: nd_reagentprop
Introduction
Interactions with Other Chado Modules
Key Ontologies
Stock Relationship Ontology
Loading Data
There are, as yet, no standard flat file formats or loading scripts to load data into this module. Custom scripts will need to be written to insert your data in the database.
Web Front Ends
See Also
- Chado Natural Diversity Module Working Group page, and the group's discussion page for background on how this module was created.
Email Threads
- Natural diversity module and phenotype cvterm values
- Use Cases on the simplified natural diversity module
- nd naming
Use Cases
ER Diagram
A simplified schema diagram by N. Menda and R. Buels, from April. This is out of date in a few ways and will be updated shortly.
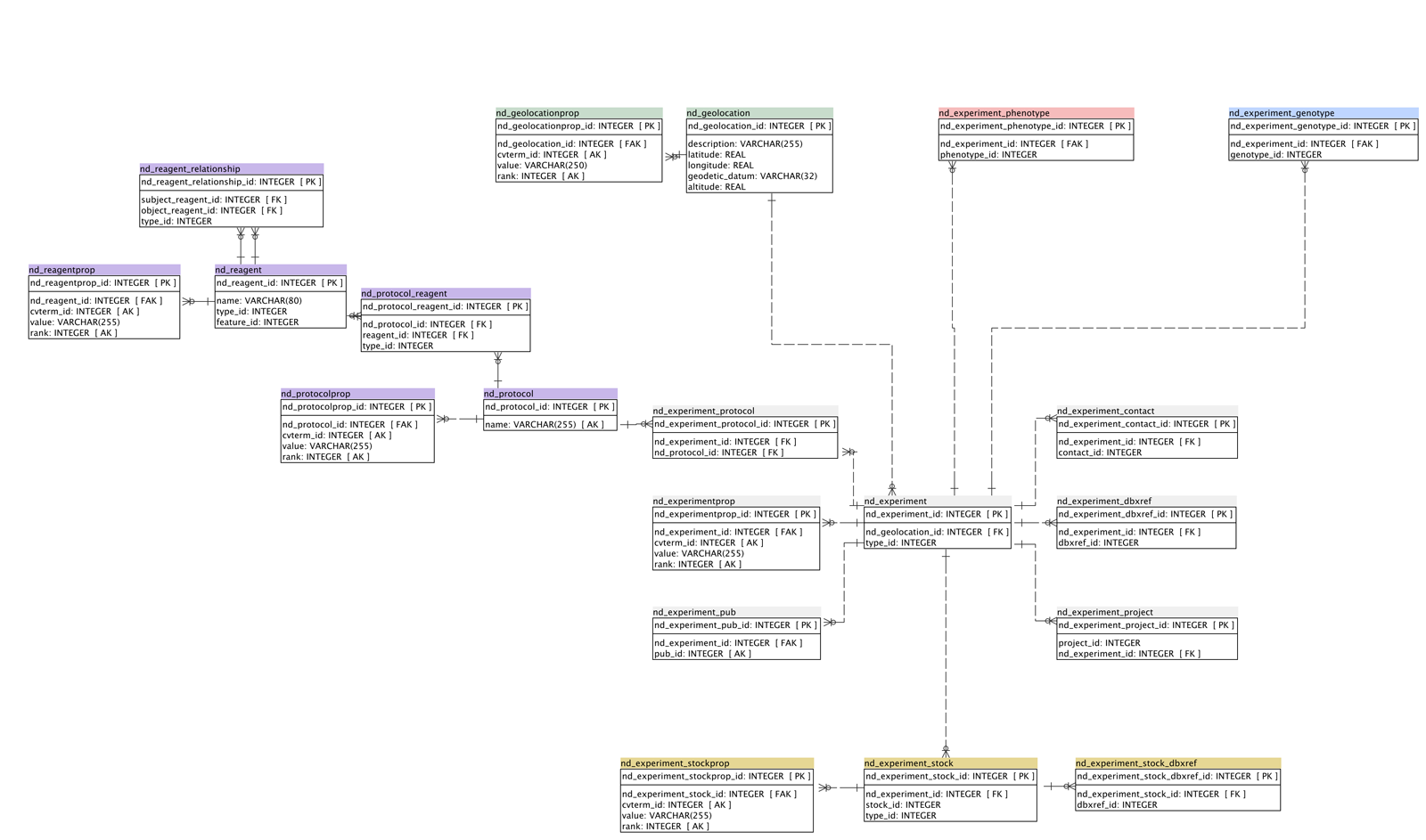
Tables
This will be populated using the process outlined in Chado Tables to Wiki.
Table: nd_experiment
This is the core table for the natural diversity module, representing each individual assay that is undertaken (nb this is usually *not* an entire experiment). Each nd_experiment should give rise to a single genotype or phenotype and be described via 1 (or more) protocols. Collections of assays that relate to each other should be linked to the same record in the project table.
Experiment.type is a cvterm that will define which records are expected for other tables. Any CV may be used but it was designed with terms such as: [phenotype_assay, genotype_assay, field_collection, cross_experiment] in mind.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_id | serial | PRIMARY KEY | |
| nd_geolocation_id | integer | NOT NULL | |
| type_id | integer | NOT NULL |
Tables referencing this one via Foreign Key Constraints:
Table: nd_experiment_contact
primary contact / submitter of these nd_experiments (nd, where assays are not submitted separately this may be better stored in project_contact).
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_contact_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | NOT NULL | |
| contact_id | integer | NOT NULL |
Table: nd_experiment_dbxref
Cross-reference experiment to accessions, images, etc
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_dbxref_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | NOT NULL | |
| dbxref_id | integer | NOT NULL |
Table: nd_experiment_genotype
Linking table: experiments to the genotypes they produce. There is a one-to-one relationship between an experiment and a genotype since each genotype record should point to one experiment. Add a new experiment_id for each genotype record.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_genotype_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | UNIQUE#1 NOT NULL | |
| genotype_id | integer | UNIQUE#1 NOT NULL |
Table: nd_experiment_phenotype
Linking table: experiments to the phenotypes they produce. There is a one-to-one relationship between an experiment and a phenotype since each phenotype record should point to one experiment. Add a new experiment_id for each phenotype record.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_phenotype_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | UNIQUE#1 NOT NULL | |
| phenotype_id | integer | UNIQUE#1 NOT NULL |
Table: nd_experiment_project
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_project_id | serial | PRIMARY KEY | |
| project_id | integer | NOT NULL | |
| nd_experiment_id | integer | NOT NULL |
Table: nd_experiment_protocol
Linking table: experiments to the protocols they involve.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_protocol_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | NOT NULL | |
| nd_protocol_id | integer | NOT NULL |
Table: nd_experiment_pub
Linking nd_experiment(s) to publication(s)
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_pub_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | UNIQUE#1 NOT NULL | |
| pub_id | integer | UNIQUE#1 NOT NULL |
Table: nd_experiment_stock
Part of a stock or a clone of a stock that is used in an experiment
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_stock_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | NOT NULL | |
| stock_id | integer | NOT NULL stock used in the extraction or the corresponding stock for the clone | |
| type_id | integer | NOT NULL |
Tables referencing this one via Foreign Key Constraints:
Table: nd_experiment_stock_dbxref
Cross-reference experiment_stock to accessions, images, etc
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_stock_dbxref_id | serial | PRIMARY KEY | |
| nd_experiment_stock_id | integer | NOT NULL | |
| dbxref_id | integer | NOT NULL |
Table: nd_experiment_stockprop
Property/value associations for experiment_stocks. This table can store the properties such as treatment
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experiment_stockprop_id | serial | PRIMARY KEY | |
| nd_experiment_stock_id | integer | UNIQUE#1 NOT NULL The experiment_stock to which the property applies. | |
| type_id | integer | UNIQUE#1 NOT NULL The name of the property as a reference to a controlled vocabulary term. | |
| value | character varying(255) | The value of the property. | |
| rank | integer | UNIQUE#1 NOT NULL The rank of the property value, if the property has an array of values. |
Table: nd_experimentprop
| FK | Name | Type | Description |
|---|---|---|---|
| nd_experimentprop_id | serial | PRIMARY KEY | |
| nd_experiment_id | integer | UNIQUE#1 NOT NULL | |
| type_id | integer | UNIQUE#1 NOT NULL | |
| value | character varying(255) | NOT NULL | |
| rank | integer | UNIQUE#1 NOT NULL |
Table: nd_geolocation
The geo-referencable location of the stock. NOTE: This entity is subject to change as a more general and possibly more OpenGIS-compliant geolocation module may be introduced into Chado.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_geolocation_id | serial | PRIMARY KEY | |
| description | character varying(255) | A textual representation of the location, if this is the original georeference. Optional if the original georeference is available in lat/long coordinates. | |
| latitude | real | The decimal latitude coordinate of the georeference, using positive and negative sign to indicate N and S, respectively. | |
| longitude | real | The decimal longitude coordinate of the georeference, using positive and negative sign to indicate E and W, respectively. | |
| geodetic_datum | character varying(32) | The geodetic system on which the geo-reference coordinates are based. For geo-references measured between 1984 and 2010, this will typically be WGS84. | |
| altitude | real | The altitude (elevation) of the location in meters. If the altitude is only known as a range, this is the average, and altitude_dev will hold half of the width of the range. |
Tables referencing this one via Foreign Key Constraints:
Table: nd_geolocationprop
Property/value associations for geolocations. This table can store the properties such as location and environment
| FK | Name | Type | Description |
|---|---|---|---|
| nd_geolocationprop_id | serial | PRIMARY KEY | |
| nd_geolocation_id | integer | UNIQUE#1 NOT NULL | |
| type_id | integer | UNIQUE#1 NOT NULL The name of the property as a reference to a controlled vocabulary term. | |
| value | character varying(250) | The value of the property. | |
| rank | integer | UNIQUE#1 NOT NULL The rank of the property value, if the property has an array of values. |
Table: nd_protocol
A protocol can be anything that is done as part of the experiment.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_protocol_id | serial | PRIMARY KEY | |
| name | character varying(255) | UNIQUE NOT NULL The protocol name. |
Tables referencing this one via Foreign Key Constraints:
Table: nd_protocol_reagent
| FK | Name | Type | Description |
|---|---|---|---|
| nd_protocol_reagent_id | serial | PRIMARY KEY | |
| nd_protocol_id | integer | NOT NULL | |
| reagent_id | integer | NOT NULL | |
| type_id | integer | NOT NULL |
Table: nd_protocolprop
Property/value associations for protocol.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_protocolprop_id | serial | PRIMARY KEY | |
| nd_protocol_id | integer | UNIQUE#1 NOT NULL The protocol to which the property applies. | |
| type_id | integer | UNIQUE#1 NOT NULL The name of the property as a reference to a controlled vocabulary term. | |
| value | character varying(255) | The value of the property. | |
| rank | integer | UNIQUE#1 NOT NULL The rank of the property value, if the property has an array of values. |
Table: nd_reagent
A reagent such as a primer, an enzyme, an adapter oligo, a linker oligo. Reagents are used in genotyping experiments, or in any other kind of experiment.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_reagent_id | serial | PRIMARY KEY | |
| name | character varying(80) | NOT NULL The name of the reagent. The name should be unique for a given type. | |
| type_id | integer | NOT NULL The type of the reagent, for example linker oligomer, or forward primer. | |
| feature_id | integer | If the reagent is a primer, the feature that it corresponds to. More generally, the corresponding feature for any reagent that has a sequence that maps to another sequence. |
Tables referencing this one via Foreign Key Constraints:
Table: nd_reagent_relationship
Relationships between reagents. Some reagents form a group. i.e., they are used all together or not at all. Examples are adapter/linker/enzyme experiment reagents.
| FK | Name | Type | Description |
|---|---|---|---|
| nd_reagent_relationship_id | serial | PRIMARY KEY | |
| subject_reagent_id | integer | NOT NULL The subject reagent in the relationship. In parent/child terminology, the subject is the child. For example, in "linkerA 3prime-overhang-linker enzymeA" linkerA is the subject, 3prime-overhand-linker is the type, and enzymeA is the object. | |
| object_reagent_id | integer | NOT NULL The object reagent in the relationship. In parent/child terminology, the object is the parent. For example, in "linkerA 3prime-overhang-linker enzymeA" linkerA is the subject, 3prime-overhand-linker is the type, and enzymeA is the object. | |
| type_id | integer | NOT NULL The type (or predicate) of the relationship. For example, in "linkerA 3prime-overhang-linker enzymeA" linkerA is the subject, 3prime-overhand-linker is the type, and enzymeA is the object. |
Table: nd_reagentprop
| FK | Name | Type | Description |
|---|---|---|---|
| nd_reagentprop_id | serial | PRIMARY KEY | |
| nd_reagent_id | integer | UNIQUE#1 NOT NULL | |
| type_id | integer | UNIQUE#1 NOT NULL | |
| value | character varying(255) | ||
| rank | integer | UNIQUE#1 NOT NULL |