JBrowse
The 2009 GMOD survey focuses on genome and comparative genomics visualization. If you use any of GMOD's visualization components (such as JBrowse), or if you have an interest in the topic, then you are strongly encouraged to take the survey before it closes on Friday, September 25. You may also win some GMOD gear.
JBrowse is a genome browser with an AJAX-based interface. JBrowse renders most tracks using client side JavaScript and JSON as its data transfer format. JBrowse is currently under development by the Ian Holmes Lab at UC Berkeley. It is expected to have a production release in 2009. Most development has been done by Mitch Skinner. JBrowse was formerly known as GBrowse 3. However, this name was changed to JBrowse to avoid confusion: JBrowse is an alternative to GBrowse, not a successor.
Contents
Demo
Requirements
- From CPAN:
- BioPerl 1.6
- JSON
- JSON::XS (optional, for speed)
- libpng
Documentation
- JBrowse Home Page
- JBrowse Quick Tutorial - JBrowse administration.
Presentations
- Image:GBrowse3GMOD2008.pdf - Presentation by Ian Holmes at July 2008 GMOD Meeting. See the talk's summary for more.
- JBrowse - Presentation by Mitch Skinner at the January 2009 GMOD Meeting. See the talk's summary for more.
- Media:Aug2009JBrowse.pdf - Presentation by Ian Holmes at the August 2009 GMOD Meeting.
Downloads
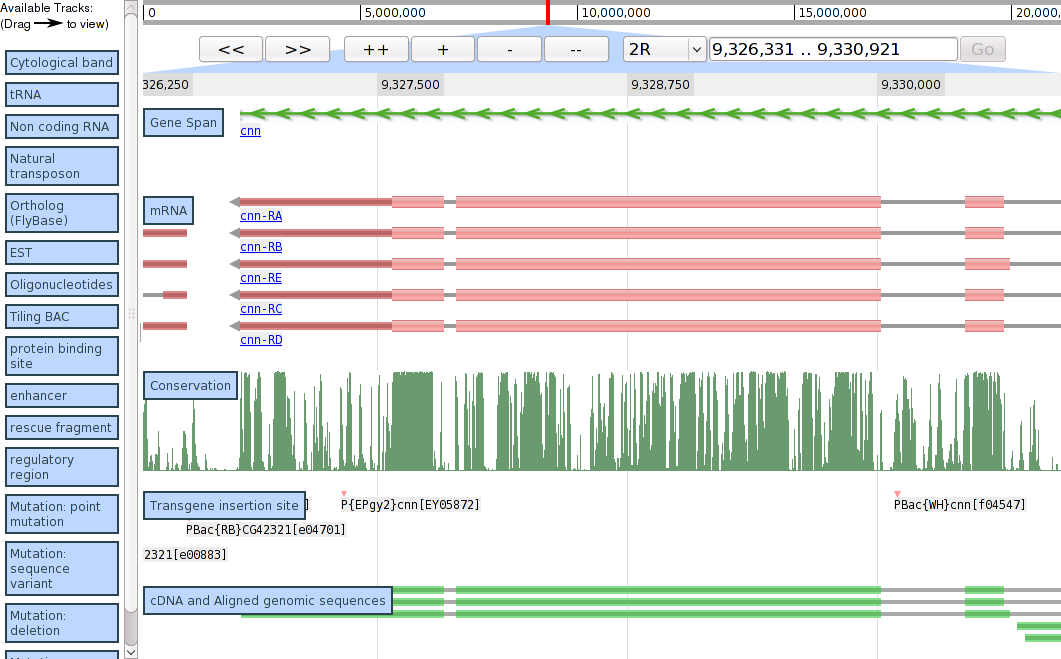
Screenshots
Demo screenshot from 2009/03/24, showing region of Drosophila melanogaster 2R.
Mailing List
- gmod-ajax@lists.sourceforge.net: Discussion of development, installation problems, etc.
- Subscribe to gmod-ajax
- gmod-ajax archives