JBrowse
| JBrowse Session 2011 GMOD Spring Training |

|
Contents
Prerequisites
These have already been set up on the VM image.
Perl:
- BioPerl 1.6
- JSON
- JSON::XS (optional, for speed)
- PerlIO::gzip
- Heap::Simple
- Heap::Simple::XS
- Devel::Size
System packages:
- libpng12-0
- libpng12-dev
Optional, for BAM files:
- samtools, and its dependency libncurses5-dev
- perl module: Bio::DB::SAM
And this is how they were installed: (don't do this)
$ sudo apt-get install git-core libpng12-0 libpng12-dev libncurses5-dev $ cd ~/Documents/Software $ wget http://sourceforge.net/projects/samtools/files/samtools/0.1.7/samtools-0.1.7a.tar.bz2 $ tar xjf samtools-0.1.7a.tar.bz2 $ cd samtools-0.1.7a/ $ make $ sudo cpan cpan[1]> install Bio::DB::Das::Chado Bio::DB::Sam JSON JSON::XS PerlIO::gzip Heap::Simple Heap::Simple::XS Devel::Size
Also: make sure you can Copy/paste from wiki.
Shell tricks:
- Tab completion
- History
- History search
JBrowse Introduction
How and why JBrowse is different from most other web-based genome browsers, including GBrowse.
More detail: paper
Media:JBrowse_GMOD_Meeting_2011.pdf
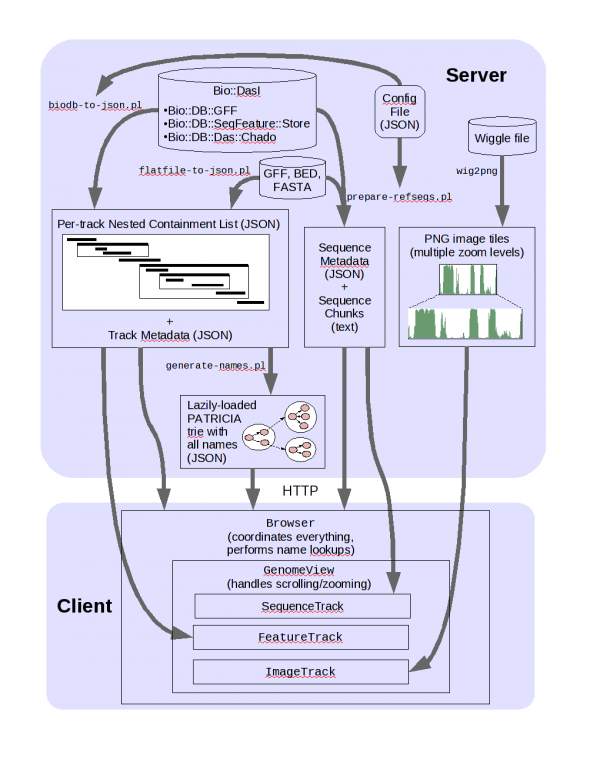
JBrowse arch
Setting up JBrowse
Getting JBrowse
- Install git (This has already been done in the VMware image.)
$ sudo apt-get install git-core
- prepare a directory for JBrowse
$ cd /var/www $ sudo mkdir jbrowse $ sudo chown gmod.gmod jbrowse
- download it from github
$ git clone git://github.com/jbrowse/jbrowse.git jbrowse
(or alternately)
$ git clone https://github.com/jbrowse/jbrowse.git jbrowse
Starting Point
Visit in web browser: http://localhost/jbrowse/
You should see just a blank white page.
Basic Steps
Setting up a JBrowse instance with feature data goes in three basic steps:
- Specify reference sequences
- Load feature data
- Collect feature names
Data from a database
Here, we'll use the Chado adapter; other common database adapters are Bio::DB::SeqFeature::Store and Bio::DB::GFF.
Starting config file: ~/Documents/Data/jbrowse/first-config.json
{
"description": "Pythium",
"db_adaptor": "Bio::DB::Das::Chado",
"db_args": { "-dsn": "dbi:Pg:dbname=chado",
"-user": "gmod",
"-pass": ""},
...
Specify reference sequences
The first script to run is bin/prepare-refseqs.pl; that script is the way you tell JBrowse about what your reference sequences are. Running bin/prepare-refseqs.pl also sets up the "DNA" track.
Run this from within the /var/www/jbrowse directory (you could run it elsewhere, but you'd have to explicitly specify the location of the data directory on the command line).
$ cd /var/www/jbrowse $ bin/prepare-refseqs.pl --conf ~/Documents/Data/jbrowse/first-config.json \ --refs scf1117875582023
Visit in web browser: you should new see the JBrowse UI (and if you zoom all the way in, some sequence)
Load Feature Data
Next, we'll use biodb-to-json.pl to get feature data out of the database and turn it into JSON data that the web browser can use.
Add a basic track definition; this will tell biodb-to-json.pl what features to put into the track, and how the track should look:
<javascript>...
"TRACK DEFAULTS": {
"class": "feature"
},
"tracks": [
{
"track": "gene",
"key": "Gene",
"feature": ["gene"],
"autocomplete": "all",
"class": "feature2",
"urlTemplate": "http://www.google.com/search?q={name}"
}
]
}</javascript>
track specifies the track identifier (a unique name for the track, for the software to use). This should be just letters and numbers and - and _ characters; using other characters makes things less convenient.
key specifies a human-friendly name for the track, which can use any characters you want.
feature gives a list of feature types to include in the track.
autocomplete including this setting makes the features in the track searchable.
urltemplate specifies a URL pattern that you can use to link genomic features to specific web pages.
class specifies the CSS class that describes how the feature should look. The classes are specified in the genome.css file:
$ less genome.css
For this particular track, I've specified the "feature2" class which looks like this in the CSS file:
<javascript>.plus-feature2, .minus-feature2 {
position:absolute; height: 15px; background-repeat: repeat-x; cursor: pointer; min-width: 1px; z-index: 10;
}
.plus-feature2 { background-image: url('img/plus-herringbone16.png'); }
.minus-feature2 { background-image: url('img/minus-herringbone16.png'); }</javascript>
Run the bin/biodb-to-json.pl script with this config file to set up this track:
$ bin/biodb-to-json.pl --conf ~/Documents/Data/jbrowse/first-config.json
(visit in web browser: you should see a new gene track)
More complex track
Now we'll add a second track; this one will have subfeatures. This snippet is from: ~/Documents/Data/jbrowse/second-config.json
<javascript>...
{
"track": "match",
"key": "Matches",
"feature": ["match"],
"autocomplete": "all",
"subfeatures": true,
"class": "generic_parent",
"subfeature_classes": {
"match_part": "match_part"
},
"clientConfig": {
"subfeatureScale": 20
}
}
...</javascript>
$ bin/biodb-to-json.pl --conf ~/Documents/Data/jbrowse/second-config.json
(visit in web browser: you should see a new track, which has subfeatures if you're zoomed in far enough)
Collect feature names
When you generate JSON for a track, if you specify "autocomplete" then a listing of all of the names/IDs from that track (along with the locations of the corresponding features) will also be generated.
The bin/generate-names.pl script collects those lists of names from all the tracks and combines them into one big tree that the client uses to search.
$ bin/generate-names.pl -v
Visit in web browser, search for feature name: e.g.,
- maker-scf1117875582023-snap-gene-0.3
Data from flat files
We're going to recreate a JBrowse instance from a different data source: flat files.
First, wipe the slate clean by removing the data directory:
$ rm -r data
If you visit your JBrowse instance in a web browser, you'll see a blank screen again
Sequences
To import sequence data from a fasta file into a JBrowse instance, use prepare-refseqs.pl with the --fasta argument:
$ bin/prepare-refseqs.pl --fasta ~/Documents/Data/jbrowse/scf1117875582023.fasta
Visit in web browser; you should see a second reference sequence.
Features
To get feature data from flat files into JBrowse, use flatfile-to-json.pl. We'll use some more of the data from the MAKER session:
$ bin/flatfile-to-json.pl \
--gff /home/gmod/Documents/Data/maker/example2_pyu/finished.maker.output/gff/scf1117875582023.gff \
--type match --getSubs --tracklabel "gff_match" --key "GFF match" \
--cssclass generic_parent --subfeatureClasses '{"match_part": "generic_part_a"}'
Visit in web browser; you should see a new "GFF match" track.
BAM data
To incorporate data from a BAM source:
$ bin/bam-to-json.pl \
--bam ~/Documents/Data/jbrowse/simulated-sorted.bam \
--tracklabel BAM_data --key "BAM Data"
Quantitative data
JBrowse can also display quantitative data in the wiggle format. JBrowse processes wiggle files with a C++ program, which you have to compile:
$ make
Now you can process the wiggle file:
$ bin/wig-to-json.pl --wig ~/Documents/Data/jbrowse/pyu.wig \
--tracklabel "coverage_wig" --key "Wiggle Coverage" --min 0 --max 50
Visit in web browser
Common Problems
- JSON syntax errors
Other links
- Config file ref: http://jbrowse.org/code/jbrowse-master/docs/config.html
- DIV test: http://jbrowse.org/test/boatdiv/boat.html
Evaluation
Please give us your comments on this session. We will ask for your feedback on each session and the course as a whole on the last day. Your comments will help guide the direction and content of future GMOD training and outreach efforts.
| Available on platform | web + |
| Has URL | http://jbrowse.org/install/ +, http://jbrowse.org +, http://twitter.com/usejbrowse +, http://github.com/GMOD/jbrowse +, http://jbrowse.org/demos +, http://icemangenome.net/%E2%80%8E +, http://genomesunzipped.org/jbrowse +, http://beetlebase.org + and http://www.medicinalgenomics.com/the-jane-ome/ + |
| Has description | Browse the genome of Ötzi the ice man + and JBrowse is a genome browser with a fully d … JBrowse is a genome browser with a fully dynamic AJAX interface, being developed as the eventual successor to GBrowse. It is very fast and scales well to large datasets. JBrowse is javascript-based and does almost all of its work directly in the user's web browser, with minimal requirements for the server.
Features[edit]
|
| Has development status | active + |
| Has input format | GFF3 +, BED +, FASTA +, WIG +, BedGraph +, Bio::DB::* +, UCSC +, Wiggle +, BigWig + and BAM + |
| Has licence | LGPL + and Artistic License 2.0 + |
| Has logo | JBrowseLogo.png + |
| Has software maturity status | mature + |
| Has support status | active + |
| Has title | JBrowse demos +, Ice Man Genome +, Genomes Unzipped: Public Personal Genomics +, BeetleBase + and The Jane-Ome, medicinal marijuana project + |
| Has topic | JBrowse + |
| Is open source | Yes + |
| Link type | download +, social media +, website +, source code +, demo server + and wild URL + |
| Release date | 2008 + |
| Tool functionality or classification | Genome visualization + |
| Written in language | Javascript + and Perl + |
| Has subobjectThis property is a special property in this wiki. | JBrowse#http://jbrowse.org/install/ +, JBrowse#http://jbrowse.org +, JBrowse#http://twitter.com/usejbrowse +, JBrowse#http://github.com/GMOD/jbrowse +, JBrowse#http://jbrowse.org/demos +, JBrowse#http://icemangenome.net/ +, JBrowse#http://genomesunzipped.org/jbrowse +, JBrowse#http://beetlebase.org + and JBrowse#http://www.medicinalgenomics.com/the-jane-ome/ + |