JBrowse
From GMOD
Status
- Mature release
- Active development
- Active support
Resources
JBrowse is a genome browser with an AJAX-based interface. JBrowse renders most tracks using client side JavaScript and JSON as its data transfer format. JBrowse is the official successor to GBrowse.
Contents
Demo
Requirements
Documentation
- JBrowseDev/Main - Thorough documentation for the most recent JBrowse release
- JBrowse Tutorial - Tutorial from 2009 GMOD Summer Schools
- JBrowse Home Page
- JBrowse Quick Tutorial - JBrowse administration.
Presentations
- Talk at the April 21, 2010 UCSC genome browser group ("genecats") meeting.
- JBrowse: A next-generation genome browser - Presentation by Ian Holmes at the August 2009 GMOD Meeting. See the talk's summary for more.
- JBrowse - Presentation by Mitch Skinner at the January 2009 GMOD Meeting. See the talk's summary for more.
- Presentation by Ian Holmes at July 2008 GMOD Meeting. See the talk's summary for more.
Downloads
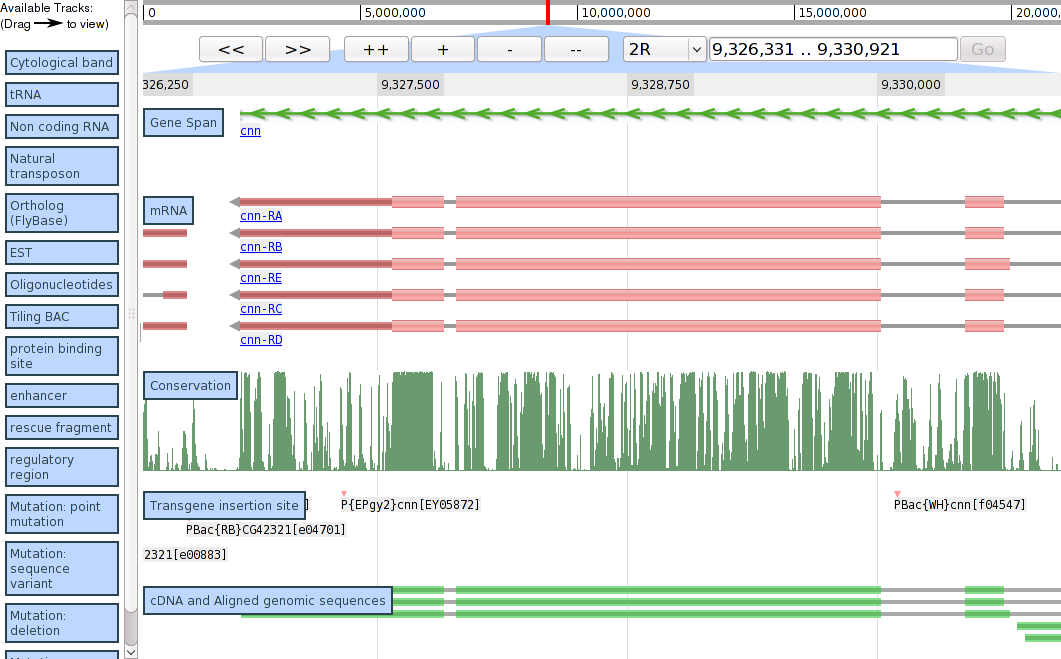
Screenshots
Demo screenshot from 2009/03/24, showing region of Drosophila melanogaster 2R.
JBrowse Development
JBrowse is an open-source project. Contributions are encouraged and welcomed. There is a developer mailing list and teleconferences on the 3rd Monday of the month at 2pm Pacific US time. Please contact Ian Holmes if you want to participate in the call.
Mailing Lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| JBrowse | gmod-ajax | JBrowse help and general questions. | Nabble (2010/05+), Sourceforge |
| jbrowse-dev | JBrowse development discussions. | Nabble (2011/08+) |