Difference between revisions of "JBrowse"
From GMOD
RobertBuels (Talk | contribs) (count the equals signs) |
RobertBuels (Talk | contribs) |
||
| Line 1: | Line 1: | ||
{{ComponentBox|{{JBrowseResourcesBoxItem}}|<!--{{ComponentBoxEventsHeader}}||{{GMODAmericas2011BoxItem|2011 GMOD Spring Training|GMOD Spring Training|March 8-12}}-->|| | | |}} | {{ComponentBox|{{JBrowseResourcesBoxItem}}|<!--{{ComponentBoxEventsHeader}}||{{GMODAmericas2011BoxItem|2011 GMOD Spring Training|GMOD Spring Training|March 8-12}}-->|| | | |}} | ||
| − | JBrowse is a genome browser with | + | JBrowse is a genome browser with a fully dynamic [[Glossary#AJAX|AJAX]] interface. JBrowse renders most tracks using client-side JavaScript, with [[Glossary#JSON|JSON]] as its data transfer format. JBrowse is being developed as the official successor to [[GBrowse]]. |
= Demo = | = Demo = | ||
[http://jbrowse.org/genomes/dmel/ ''D. melanogaster''] | [http://jbrowse.org/genomes/dmel/ ''D. melanogaster''] | ||
| + | |||
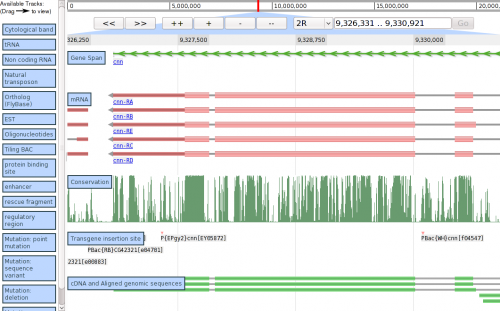
| + | [[Image:JBrowseShot2.png|border|500px|Demo screenshot from 2009/03/24, showing region of ''Drosophila melanogaster'' 2R.]] | ||
=Installation= | =Installation= | ||
| Line 49: | Line 51: | ||
[http://github.com/jbrowse From GitHub]. | [http://github.com/jbrowse From GitHub]. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
= JBrowse Development = | = JBrowse Development = | ||
Revision as of 21:23, 26 October 2011
Status
- Mature release
- Active development
- Active support
Resources
JBrowse is a genome browser with a fully dynamic AJAX interface. JBrowse renders most tracks using client-side JavaScript, with JSON as its data transfer format. JBrowse is being developed as the official successor to GBrowse.
Contents
Demo
Installation
See Installing JBrowse on Mac OS X and Linux for step-by-step installation instructions.
Prerequisites
Usage Notes
- General Usage, including information about:
- The JBrowse user interface
- Creating sequence tracks (e.g. from FASTA files, or a Chado database)
- Creating feature tracks (e.g. from BED or GFF files, a Chado database, or the UCSC genome browser)
- Creating image tracks (e.g. from WIG files)
- Making features searchable by name
- Removing tracks
- Additional topics:
Presentations
- Talk at the April 21, 2010 UCSC genome browser group ("genecats") meeting.
- JBrowse: A next-generation genome browser - Presentation by Ian Holmes at the August 2009 GMOD Meeting. See the talk's summary for more.
- JBrowse - Presentation by Mitch Skinner at the January 2009 GMOD Meeting. See the talk's summary for more.
- Presentation by Ian Holmes at July 2008 GMOD Meeting. See the talk's summary for more.
See also
- JBrowse Home Page
- JBrowse Quick Tutorial - JBrowse administration.
- JBrowseDev/Upcoming Future JBrowse features
Downloads
JBrowse Development
JBrowse is an open-source project. Contributions are encouraged and welcomed. There is a developer mailing list and teleconferences on the 3rd Monday of the month at 2pm Pacific US time. Please contact Ian Holmes if you want to participate in the call.
Contact
- Mailing lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| JBrowse | gmod-ajax | JBrowse help and general questions. | Nabble (2010/05+), Sourceforge |
| jbrowse-dev | JBrowse development discussions. | Nabble (2011/08+) |
- Original author
- Mitch Skinner