Difference between revisions of "GBrowse syn Tutorial 2012"
m (Adding AMI info) |
|||
| (18 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| − | + | <div class=emphasisbox style="font-size:36pt"> | |
| + | If you are 2013 GMOD Summer School, you are on the wrong tutorial.<br> | ||
| + | Use [[GBrowse_syn_Tutorial_2013]]. | ||
| + | </div> | ||
| − | |||
| + | This tutorial on [[GBrowse syn]] was taught by [[User:Mckays|Sheldon McKay]] as part of the [[2013 GMOD Summer School]]. | ||
| + | |||
| + | {{Template:AMI Summer School day 4}} | ||
| + | * If you are not using the Amazon EC2 instance, the system paths may vary from those described below. | ||
| + | | ||
[[GBrowse_syn|GBrowse_syn]] is a [[GBrowse|GBrowse]]-based [[Synteny|synteny]] browser designed to display multiple genomes, with a central reference species compared to two or more additional species. It is included with the standard GBrowse package (version 1.69 and later). | [[GBrowse_syn|GBrowse_syn]] is a [[GBrowse|GBrowse]]-based [[Synteny|synteny]] browser designed to display multiple genomes, with a central reference species compared to two or more additional species. It is included with the standard GBrowse package (version 1.69 and later). | ||
| Line 9: | Line 16: | ||
[[File:Gbrowse_syn.png|thumb|none|600px|GBrowse_syn at The Arabidopsis Information Resource (TAIR)]] | [[File:Gbrowse_syn.png|thumb|none|600px|GBrowse_syn at The Arabidopsis Information Resource (TAIR)]] | ||
| − | Working examples of GBrowse_syn can be seen at [http://www.arabidopsis.org/cgi-bin/gbrowse_syn/arabidopsis/?name=Chr1%3A8367000..8370501;search_src=thaliana TAIR] and [http:// | + | Working examples of GBrowse_syn can be seen at [http://www.arabidopsis.org/cgi-bin/gbrowse_syn/arabidopsis/?name=Chr1%3A8367000..8370501;search_src=thaliana TAIR] and [http://mckay.cshl.edu/seq/gbrowse_syn/compara?search_src=Cele;name=X:1050001..1150000 WormBase]. |
| Line 34: | Line 41: | ||
* The example we will use is a two-species comparison of rice (''Oryza sativa'') and one of its wild relatives* | * The example we will use is a two-species comparison of rice (''Oryza sativa'') and one of its wild relatives* | ||
| − | <div class="zero small"> | + | <div class="zero small">*Data courtesy of Bonnie Hurwitz</div> |
</div> | </div> | ||
| Line 40: | Line 47: | ||
The alignment, or joining database will contain the sequence alignments between the two rice species. It will be in a [[MySQL|MySQL]] database. | The alignment, or joining database will contain the sequence alignments between the two rice species. It will be in a [[MySQL|MySQL]] database. | ||
| + | If mysql is not installed on your server, you can install it as follows: | ||
| + | '''Debian-based Linux distributions:''' | ||
| + | $ sudo apt-get update | ||
| + | $ sudo apt-get install mysql-client | ||
| + | $ sudo apt-get install mysql-servery | ||
| + | |||
| + | '''RedHat-based Linux distributions''' | ||
| + | $ sudo yum install mysql | ||
| + | <div class="emphasisbox"> | ||
| + | Note: You set the mysql root password at the time of installation. Use 'gbsyndemo' or else be sure to remember the password for use later. | ||
| + | </div> | ||
1) Create a MySQL database to hold the alignment data | 1) Create a MySQL database to hold the alignment data | ||
$ <span class="enter">mysql -u root -p</span> | $ <span class="enter">mysql -u root -p</span> | ||
Enter password: <span class="enter">****************</span> | Enter password: <span class="enter">****************</span> | ||
| − | + | <pre> | |
| + | Welcome to the MySQL monitor. Commands end with ; or \g. | ||
Your MySQL connection id is 37 | Your MySQL connection id is 37 | ||
Server version: 5.1.37-1ubuntu5.1 (Ubuntu) | Server version: 5.1.37-1ubuntu5.1 (Ubuntu) | ||
| Line 51: | Line 70: | ||
Type 'help;' or '\h' for help. Type '\c' to clear the current input statement. | Type 'help;' or '\h' for help. Type '\c' to clear the current input statement. | ||
| − | mysql | + | mysql> create database rice_synteny; |
Query OK, 1 row affected (0.00 sec) | Query OK, 1 row affected (0.00 sec) | ||
mysql> | mysql> | ||
| + | </pre> | ||
2) Give read-only (SELECT privileges in SQL) to the default apache user <tt>www-data</tt>. We can do this for all of the MySQL databases, since they are all for web applications | 2) Give read-only (SELECT privileges in SQL) to the default apache user <tt>www-data</tt>. We can do this for all of the MySQL databases, since they are all for web applications | ||
| Line 63: | Line 83: | ||
mysql> <span class="enter">quit</span> | mysql> <span class="enter">quit</span> | ||
| − | 3) | + | 3) Load the sample alignment database. You need to have root-level access (be a sudoer) for some of the steps below. |
| − | $ <span class="enter">cd /var/ | + | $ <span class="enter">cd /data/var/lib/gbrowse2/databases/gbrowse_syn/alignments</span> |
| − | + | ||
Have a look at the first few lines of the data: | Have a look at the first few lines of the data: | ||
| − | $ <span class="enter"> | + | $ <span class="enter">zcat rice.aln.gz | head -20</span> |
| + | <pre> | ||
CLUSTAL W(1.81) multiple sequence alignment W(1.81) | CLUSTAL W(1.81) multiple sequence alignment W(1.81) | ||
| Line 91: | Line 111: | ||
rice-3(+)/16598648-16600199 tggagcctccccttctagctcgatcacgctctgctcttccgcttggaggctggcaaaact | rice-3(+)/16598648-16600199 tggagcctccccttctagctcgatcacgctctgctcttccgcttggaggctggcaaaact | ||
wild_rice-3(+)/14467855-14469373 tggagcctccccttctagctcgatcgcgctctgctcttccgcttggaggctggcaaaact | wild_rice-3(+)/14467855-14469373 tggagcctccccttctagctcgatcgcgctctgctcttccgcttggaggctggcaaaact | ||
| + | </pre> | ||
The format is CLUSTALW. This is a formatting convention; it does not mean CLUSTALW was used to generate the alignment data. See [[#Further Reading|Further Reading]] below for more information on data loading and the meta-data in the sequence names | The format is CLUSTALW. This is a formatting convention; it does not mean CLUSTALW was used to generate the alignment data. See [[#Further Reading|Further Reading]] below for more information on data loading and the meta-data in the sequence names | ||
| − | + | Load the database with the script <tt>gbrowse_syn_load_alignments_msa.pl</tt>, which is automatically installed along with GBrowse. See the [GBrowse_syn scripts] page for details on the options for the script. | |
| − | $ <span class="enter">gbrowse_syn_load_alignments_msa.pl -u root -p | + | $ <span class="enter">zcat rice.aln.gz | gbrowse_syn_load_alignments_msa.pl -u root -p ******* -d rice_synteny -c -v -</span> |
There are 1800 alignment blocks, so this will take a little while to run. | There are 1800 alignment blocks, so this will take a little while to run. | ||
| Line 102: | Line 123: | ||
===Setting up the Configuration Files=== | ===Setting up the Configuration Files=== | ||
<div class="emphasisbox"> | <div class="emphasisbox"> | ||
| − | * The configuration files required for this data source are pre-installed with [[GBrowse]], in <tt>/etc/gbrowse2/synteny/</tt>. | + | * The configuration files required for this data source are pre-installed with [[GBrowse]], in <tt>/data/etc/gbrowse2/synteny/</tt>. |
* There are config files for two species, <tt>rice_synteny.conf</tt> and <tt>wild_rice_synteny.conf</tt>, and the joining config file, <tt>oryza.synconf</tt>. The latter file has been disabled by appending a '.disabled' extension to the file name. | * There are config files for two species, <tt>rice_synteny.conf</tt> and <tt>wild_rice_synteny.conf</tt>, and the joining config file, <tt>oryza.synconf</tt>. The latter file has been disabled by appending a '.disabled' extension to the file name. | ||
</div> | </div> | ||
| Line 188: | Line 209: | ||
2) Renaming the configuration file | 2) Renaming the configuration file | ||
| − | $ <span class="enter">cd /etc/gbrowse2/synteny</span> | + | $ <span class="enter">cd /data/etc/gbrowse2/synteny</span> |
$ <span class="enter">sudo mv oryza.synconf.disabled oryza.synconf</span> | $ <span class="enter">sudo mv oryza.synconf.disabled oryza.synconf</span> | ||
| − | 3) Point your browser to <nowiki>http://</nowiki>{{Template:AWSurl}}/cgi-bin/gb2/gbrowse_syn/oryza. You should see: | + | 3) Point your browser to <nowiki>http://</nowiki>{{Template:AWSurl}}/cgi-bin/gb2/gbrowse_syn/oryza (or your own URL if you are not using the Amazon EC2 instance). You should see: |
[[Image:GBrowse_synWe_made_it1.png|thumb|none|800px]] | [[Image:GBrowse_synWe_made_it1.png|thumb|none|800px]] | ||
| Line 204: | Line 225: | ||
[[File:Gbrowse_synBubble1.png]] | [[File:Gbrowse_synBubble1.png]] | ||
* Click on one of the bold blue highlighted section titles. This takes you to a contextual help page on the GMOD wiki. | * Click on one of the bold blue highlighted section titles. This takes you to a contextual help page on the GMOD wiki. | ||
| − | * Click and drag on the overview panel. This will trigger rubber band selection to recenter or resize the displayed image | + | <!--* Click and drag on the overview panel. This will trigger rubber band selection to recenter or resize the displayed image--> |
===Speeding up the Browser=== | ===Speeding up the Browser=== | ||
| Line 266: | Line 287: | ||
$ <span class="enter">wget ftp://ftp.gmod.org/pub/gmod/GBrowse_syn/orthocluster.tar.gz</span> | $ <span class="enter">wget ftp://ftp.gmod.org/pub/gmod/GBrowse_syn/orthocluster.tar.gz</span> | ||
$ <span class="enter">tar zxf orthocluster.tar.gz</span> | $ <span class="enter">tar zxf orthocluster.tar.gz</span> | ||
| + | $ tar zxvf orthocluster.tar.gz | ||
| + | ORTHOCLUSTER/ | ||
| + | ORTHOCLUSTER/conf/ | ||
| + | ORTHOCLUSTER/gff/ | ||
| + | ORTHOCLUSTER/orthocluster.txt | ||
| + | ORTHOCLUSTER/gff/bri.gff | ||
| + | ORTHOCLUSTER/gff/ele.gff | ||
| + | ORTHOCLUSTER/gff/ppa.gff | ||
| + | ORTHOCLUSTER/conf/bri.conf | ||
| + | ORTHOCLUSTER/conf/ele.conf | ||
| + | ORTHOCLUSTER/conf/orthocluster.synconf | ||
| + | ORTHOCLUSTER/conf/ppa.conf | ||
| − | |||
| − | + | In the <tt>conf</tt> directory, there are configuration files for the joining database and each of the three species. Note that the annotation data are in old-school GFF3 format: | |
| − | + | <span class=enter>head -20 bri.gff</span> | |
| − | + | $ head -20 ele.gff | |
| − | + | ##gff-version 2 | |
| − | + | ##sequence-region I 1 15072421 | |
| − | + | ##sequence-region II 1 15279324 | |
| − | + | ##sequence-region III 1 13783685 | |
| − | + | ##sequence-region IV 1 17493784 | |
| − | + | ##sequence-region V 1 20924143 | |
| − | + | ##sequence-region X 1 17718854 | |
| − | + | I curated coding_exon 11641 11689 . + 0 CDS "Y74C9A.2" | |
| − | + | I curated coding_exon 14951 15160 . + 2 CDS "Y74C9A.2" | |
| − | + | I curated coding_exon 16473 16585 . + 2 CDS "Y74C9A.2" | |
| − | + | I curated coding_exon 43733 43961 . + 0 CDS "Y74C9A.1" | |
| − | + | I curated coding_exon 44030 44234 . + 2 CDS "Y74C9A.1" | |
| − | + | I curated coding_exon 44281 44324 . + 1 CDS "Y74C9A.1" | |
| − | + | I curated coding_exon 44372 44468 . + 2 CDS "Y74C9A.1" | |
| + | I curated coding_exon 44521 44677 . + 1 CDS "Y74C9A.1" | ||
| + | I curated coding_exon 47472 47610 . + 0 CDS "Y48G1C.12" | ||
| + | I curated coding_exon 47696 47858 . + 2 CDS "Y48G1C.12" | ||
| + | I curated coding_exon 48348 48530 . + 1 CDS "Y48G1C.12" | ||
| + | I curated coding_exon 49251 49416 . + 1 CDS "Y48G1C.12" | ||
| − | + | We know that there are only split CDS features: | |
| + | <span class=enter>cut -f3 ele.gff | sort -u</span> | ||
| − | + | We can convert this simple GFF to GFF3 with a perl one-liner: | |
| − | + | <span class=enter>perl -pe 's/version 2/version 3/;s/CDS /Parent=/;s/coding_exon/CDS/' ele.gff >ele.gff3</span> | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | Then load into a Bio::DB::SeqFeature::Store database | |
| + | <span class=enter>mysql -uroot -pgbsyndemo -e 'create database ele'</span> | ||
| + | <span class=enter>bp_seqfeature_load -u root -p gbsyndemo -d ele -c -f ele.gff3</span> | ||
| − | |||
<pre> | <pre> | ||
| + | |||
[GENERAL] | [GENERAL] | ||
description = C. briggsae | description = C. briggsae | ||
Latest revision as of 17:23, 22 July 2013
If you are 2013 GMOD Summer School, you are on the wrong tutorial.
Use GBrowse_syn_Tutorial_2013.
This tutorial on GBrowse syn was taught by Sheldon McKay as part of the 2013 GMOD Summer School.
To follow along with the tutorial, you will need to use AMI ID: ami-5bab1c32, name: GMOD 2012 day 4 start, available in the US East (N. Virginia) region. See the GMOD Cloud Tutorial for information on how to get this AMI.
- If you are not using the Amazon EC2 instance, the system paths may vary from those described below.
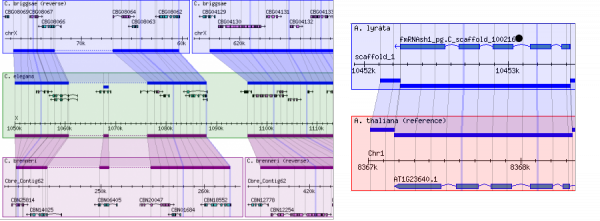
GBrowse_syn is a GBrowse-based synteny browser designed to display multiple genomes, with a central reference species compared to two or more additional species. It is included with the standard GBrowse package (version 1.69 and later).
Working examples of GBrowse_syn can be seen at TAIR and WormBase.
Contents
GBrowse_syn Introduction
Introductory talk on GBrowse_syn
Installing GBrowse_syn
GBrowse_syn is part of the GBrowse 2.0 package and was pre-installed when you went through the GBrowse 2.0 installation.
Getting Started
Point your browser to http://ec2-##-##-##-##.compute-1.amazonaws.com/cgi-bin/gb2/gbrowse_syn
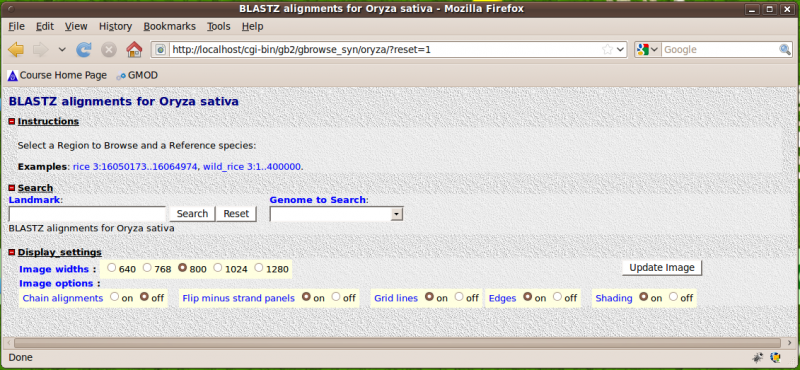
This is the welcome screen you should see after installing a new copy of GBrowse_syn with no configured data sources. It contains instructions on how to set up the example data source provided with the distribution.
Setting up the sample data
- Sample data and configuration information for GBrowse_syn come pre-packaged with GBrowse.
- The example we will use is a two-species comparison of rice (Oryza sativa) and one of its wild relatives*
Setting up the Alignment Database
The alignment, or joining database will contain the sequence alignments between the two rice species. It will be in a MySQL database. If mysql is not installed on your server, you can install it as follows:
Debian-based Linux distributions:
$ sudo apt-get update $ sudo apt-get install mysql-client $ sudo apt-get install mysql-servery
RedHat-based Linux distributions
$ sudo yum install mysql
Note: You set the mysql root password at the time of installation. Use 'gbsyndemo' or else be sure to remember the password for use later.
1) Create a MySQL database to hold the alignment data
$ mysql -u root -p Enter password: ****************
Welcome to the MySQL monitor. Commands end with ; or \g. Your MySQL connection id is 37 Server version: 5.1.37-1ubuntu5.1 (Ubuntu) Type 'help;' or '\h' for help. Type '\c' to clear the current input statement. mysql> create database rice_synteny; Query OK, 1 row affected (0.00 sec) mysql>
2) Give read-only (SELECT privileges in SQL) to the default apache user www-data. We can do this for all of the MySQL databases, since they are all for web applications
mysql> GRANT SELECT on *.* TO 'www-data'@'localhost';
Query OK, 0 rows affected (0.00 sec)
mysql> quit
3) Load the sample alignment database. You need to have root-level access (be a sudoer) for some of the steps below.
$ cd /data/var/lib/gbrowse2/databases/gbrowse_syn/alignments
Have a look at the first few lines of the data:
$ zcat rice.aln.gz | head -20
CLUSTAL W(1.81) multiple sequence alignment W(1.81)
rice-3(+)/16598648-16600199 ggaggccggccgtctgccatgcgtgagccagacggggcgggccggagacaggccacgtgg
wild_rice-3(+)/14467855-14469373 gggggccgg------------------------------------agacaggccacgtgg
** ****** ***************
rice-3(+)/16598648-16600199 ccctgccccgggctgttgacccactggcacccctgtcccgggttgtcgccctcctttccc
wild_rice-3(+)/14467855-14469373 ccctgccccgggctgttgacccactggcacccctgtcccgggttgtcgccctcctttccc
************************************************************
rice-3(+)/16598648-16600199 cgccatgctctaagtttgctcctcttctcgaacttctctctttgattcttcacgtcctct
wild_rice-3(+)/14467855-14469373 cgccatgctctaagtttgctcctcttctcgaacttctctctttgattcttcacgtcctct
************************************************************
rice-3(+)/16598648-16600199 tggagcctccccttctagctcgatcacgctctgctcttccgcttggaggctggcaaaact
wild_rice-3(+)/14467855-14469373 tggagcctccccttctagctcgatcgcgctctgctcttccgcttggaggctggcaaaact
The format is CLUSTALW. This is a formatting convention; it does not mean CLUSTALW was used to generate the alignment data. See Further Reading below for more information on data loading and the meta-data in the sequence names
Load the database with the script gbrowse_syn_load_alignments_msa.pl, which is automatically installed along with GBrowse. See the [GBrowse_syn scripts] page for details on the options for the script.
$ zcat rice.aln.gz | gbrowse_syn_load_alignments_msa.pl -u root -p ******* -d rice_synteny -c -v -
There are 1800 alignment blocks, so this will take a little while to run.
Setting up the Configuration Files
- The configuration files required for this data source are pre-installed with GBrowse, in /data/etc/gbrowse2/synteny/.
- There are config files for two species, rice_synteny.conf and wild_rice_synteny.conf, and the joining config file, oryza.synconf. The latter file has been disabled by appending a '.disabled' extension to the file name.
The joining config file, oryza.synconf:
[GENERAL]
description = BLASTZ alignments for Oryza sativa
====Sample Configuration Files====
# The synteny database
join = dbi:mysql:database=rice_synteny;host=localhost
# This option maps the relationship between the species data sources, names and descriptions
# The value for "name" (the first column) is the symbolic name that gbrowse_syn users to identify each species.
# This value is also used in two other places in the gbrowse_syn configuration:
# the species name in the "examples" directive and the species name in the .aln file
# The value for "conf. file" is the basename of the corresponding gbrowse .conf files.
# This value is also used to identify the species configuration stanzas at the bottom of the configuration file.
# name conf. file Description
source_map = rice rice_synteny "Domesic Rice (O. sativa)"
wild_rice wild_rice_synteny "Wild Rice"
tmpimages = /tmp/gbrowse2
imagewidth = 800
stylesheet = /gbrowse2/css/gbrowse_transparent.css
cache time = 1
config_extension = conf
# example searches to display
examples = rice 3:16050173..16064974
wild_rice 3:1..400000
zoom levels = 5000 10000 25000 50000 100000 200000 400000
# species-specific databases
[rice_synteny]
tracks = EG
color = blue
[wild_rice_synteny]
tracks = EG
color = red
A sample species config file, rice_synteny.conf:
[GENERAL]
description = Domestic rice chromosome 3
db_adaptor = Bio::DB::SeqFeature::Store
db_args = -adaptor memory
-dir /var/www/gbrowse2/databases/gbrowse_syn/rice
# Web site configuration info
tmpimages = /tmp/gbrowse2
[EG]
feature = gene:ensembl
glyph = gene
height = 10
bgcolor = peachpuff
fgcolor = hotpink
description = 0
label = 0
category = Transcripts
key = ensembl gene
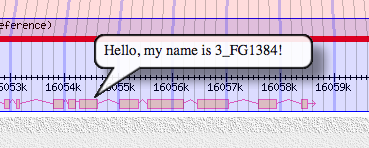
balloon hover = Hello, my name is $name!
Note: the species databases are actually using the GFF3 flat file, in-memory adapter
Activating the Oryza Data Source
1) Make sure the temporary image directory specified in the config files exists and is world-writable
$ sudo mkdir /var/www/tmp $ sudo mkdir /var/www/tmp/gbrowse2 $ sudo chmod 777 /var/www/tmp/gbrowse2
2) Renaming the configuration file
$ cd /data/etc/gbrowse2/synteny $ sudo mv oryza.synconf.disabled oryza.synconf
3) Point your browser to http://ec2-##-##-##-##.compute-1.amazonaws.com/cgi-bin/gb2/gbrowse_syn/oryza (or your own URL if you are not using the Amazon EC2 instance). You should see:
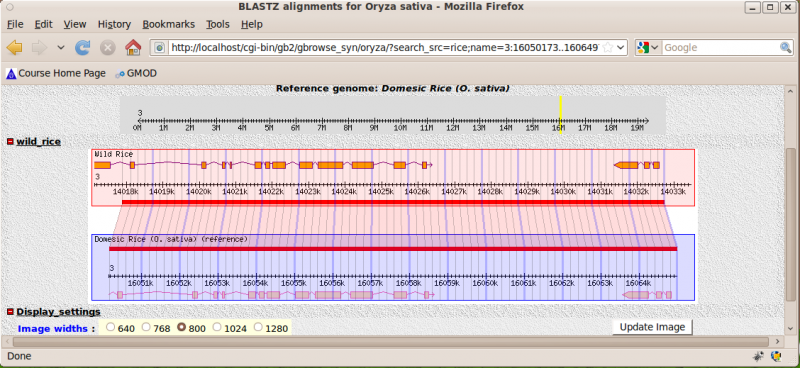
4) Click on the first example, you should (eventually) see:
5) Try out a few user interface features:
- mouse over one of the genes:
- Click on one of the bold blue highlighted section titles. This takes you to a contextual help page on the GMOD wiki.
Speeding up the Browser
You can speed up the image loading time by putting your species' GFF3 data into relational MySQL databases.
1) Create a database for each of the GFF data files (rice.gff3 and wild_rice.gff3).
$ mysql -uroot -pgbsyndemo
mysql> create database rice;
Query OK, 1 row affected (0.00 sec)
mysql> create database wild_rice;
Query OK, 1 row affected (0.00 sec)
mysql> quit
2) Populate the databases using the Loading bp_seqfeature_load.pl (pre-installed as part of BioPerl with GBrowse). This will load the GFF3 data into a MySQL relational database. Note the MySQL user will root-level privileges.
$ cd /var/www/gbrowse2/databases/gbrowse_syn/rice $ bp_seqfeature_load.pl -u root -p gbsyndemo -d rice -c -f rice.gff3 loading rice.gff3... Building object tree... 0.55s0s
Loading bulk data into database... 0.73s load time: 11.99s
$ cd ../wild_rice $ bp_seqfeature_load.pl -u root -p gbsyndemo -d wild_rice -c -f wild_rice.gff3 loading wild_rice.gff3... Building object tree... 0.55s7a
Loading bulk data into database... 0.69s load time: 12.02s
3) Modify the following stanza in the file rice_synteny.conf in cd /etc/gbrowse2/synteny/. This will convert your data source from a flat file database to a MySQL relational database.
# from
db_args = -adaptor memory
-dir /var/www/html/gbrowse/databases/gbrowse_syn/rice
# to
db_args = -dsn dbi:mysql:rice
4) repeat for wild_rice_synteny.conf
Using Non-alignment Data
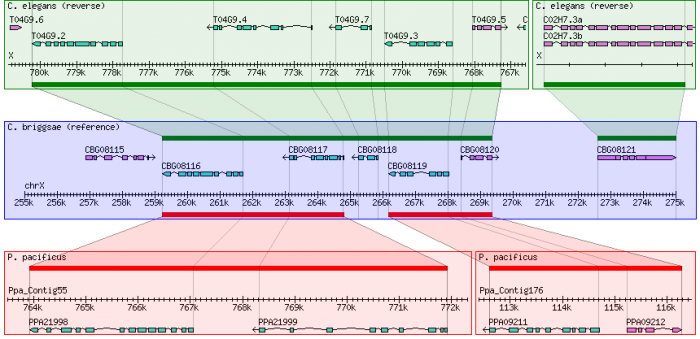
This example uses gene orthology-based synteny blocks* based created by OrthoCluster for three nematode species, C. elegans, C. briggsae and P. pacificus.
1) Download and unpack the data archive file orthocluster.tar.gz.
$ cd ~/Documents/Data/gbrowse_syn $ rm orthocluster.tar.gz $ wget ftp://ftp.gmod.org/pub/gmod/GBrowse_syn/orthocluster.tar.gz $ tar zxf orthocluster.tar.gz
$ tar zxvf orthocluster.tar.gz
ORTHOCLUSTER/ ORTHOCLUSTER/conf/ ORTHOCLUSTER/gff/ ORTHOCLUSTER/orthocluster.txt ORTHOCLUSTER/gff/bri.gff ORTHOCLUSTER/gff/ele.gff ORTHOCLUSTER/gff/ppa.gff ORTHOCLUSTER/conf/bri.conf ORTHOCLUSTER/conf/ele.conf ORTHOCLUSTER/conf/orthocluster.synconf ORTHOCLUSTER/conf/ppa.conf
In the conf directory, there are configuration files for the joining database and each of the three species. Note that the annotation data are in old-school GFF3 format:
head -20 bri.gff
$ head -20 ele.gff
##gff-version 2 ##sequence-region I 1 15072421 ##sequence-region II 1 15279324 ##sequence-region III 1 13783685 ##sequence-region IV 1 17493784 ##sequence-region V 1 20924143 ##sequence-region X 1 17718854 I curated coding_exon 11641 11689 . + 0 CDS "Y74C9A.2" I curated coding_exon 14951 15160 . + 2 CDS "Y74C9A.2" I curated coding_exon 16473 16585 . + 2 CDS "Y74C9A.2" I curated coding_exon 43733 43961 . + 0 CDS "Y74C9A.1" I curated coding_exon 44030 44234 . + 2 CDS "Y74C9A.1" I curated coding_exon 44281 44324 . + 1 CDS "Y74C9A.1" I curated coding_exon 44372 44468 . + 2 CDS "Y74C9A.1" I curated coding_exon 44521 44677 . + 1 CDS "Y74C9A.1" I curated coding_exon 47472 47610 . + 0 CDS "Y48G1C.12" I curated coding_exon 47696 47858 . + 2 CDS "Y48G1C.12" I curated coding_exon 48348 48530 . + 1 CDS "Y48G1C.12" I curated coding_exon 49251 49416 . + 1 CDS "Y48G1C.12"
We know that there are only split CDS features:
cut -f3 ele.gff | sort -u
We can convert this simple GFF to GFF3 with a perl one-liner:
perl -pe 's/version 2/version 3/;s/CDS /Parent=/;s/coding_exon/CDS/' ele.gff >ele.gff3
Then load into a Bio::DB::SeqFeature::Store database
mysql -uroot -pgbsyndemo -e 'create database ele' bp_seqfeature_load -u root -p gbsyndemo -d ele -c -f ele.gff3
[GENERAL]
description = C. briggsae
db_adaptor = Bio::DB::GFF
db_args = -dsn dbi:mysql:bri
# This is the GFF2 aggregator that assembles gene models
# from coding exon features with the same parent
aggregators = gene{coding_exon}
The gff directory contains gene annotations for each of the three species, derived from WormBase (release WS204). The files are in GFF2 format, which is why the Bio::DB::GFF adapter is required. A sample is shown here:
##gff-version 2 ##sequence-region I 1 15072421 ##sequence-region II 1 15279324 ##sequence-region III 1 13783685 ##sequence-region IV 1 17493784 ##sequence-region V 1 20924143 ##sequence-region X 1 17718854 I curated coding_exon 11641 11689 . + 0 CDS "Y74C9A.2" I curated coding_exon 14951 15160 . + 2 CDS "Y74C9A.2" I curated coding_exon 16473 16585 . + 2 CDS "Y74C9A.2" I curated coding_exon 43733 43961 . + 0 CDS "Y74C9A.1" I curated coding_exon 44030 44234 . + 2 CDS "Y74C9A.1" I curated coding_exon 44281 44324 . + 1 CDS "Y74C9A.1" I curated coding_exon 44372 44468 . + 2 CDS "Y74C9A.1" I curated coding_exon 44521 44677 . + 1 CDS "Y74C9A.1" I curated coding_exon 47472 47610 . + 0 CDS "Y48G1C.12" I curated coding_exon 47696 47858 . + 2 CDS "Y48G1C.12" I curated coding_exon 48348 48530 . + 1 CDS "Y48G1C.12" I curated coding_exon 49251 49416 . + 1 CDS "Y48G1C.12"
The file orthocluster.txt contains the synteny data. The first few lines are shown below. The first 12 fields in each row specify information about the synteny block in each species and the series of numbers are orthologous gene coordinate pairs that are used for linking orthologs with grid-lines in the GBrowse_syn display. See 'Alignment Data' under Further Reading below for more details of this loading format.
bri chrI 176154 183558 + . ppa Ppa_Contig88 27212 30786 + . 176154 27212 177594 30786 182118 27212 183558 30786 | 30786 183558 27212 182118 30786 177594 27212 176154 bri chrI 778780 799223 + . ppa Ppa_Contig88 533454 542961 - . 778780 539924 786778 542961 789497 533454 799223 538726 | 538726 799223 533454 789497 542961 786778 539924 778780 bri chrI 986150 994698 + . ppa Ppa_Contig77 29481 45600 - . 986150 37055 989649 45600 991428 29481 994698 36608 | 36608 994698 29481 991428 45600 989649 37055 986150 bri chrI 1453793 1461931 + . ppa Ppa_Contig132 156183 165414 - . 1453793 163110 1456404 165414 1456712 160849 1457637 162712 1458361 160204 1459245 160815 1459468 159346 1459854 160000 1459962 156183 1461931 159022 | 159022 1461931 156183 1459962 160000 1459854 159346 1459468 160815 1459245 160204 1458361 162712 1457637 160849 1456712 165414 1456404 163110 1453793
3) Set the $TMP environmental variable so that the database loading script knows where to put its temp files.
$ export TMP=/tmp
4) Create and load a Bio::DB:GFF database for C. elegans (ele). Use screen so that we can get the time-consuming loading script started and then use Ctrl-A D to set the screen running in the background and move on to other steps.
$ cd ORTHOCLUSTER/gff $ mysql -uroot -pgbsyndemo -e 'create database ele' $ screen bp_fast_load_gff.pl -u root -p gbsyndemo -d ele -c ele.gff
5) Repeat step 4 for the other two species (bri and ppa).
6) Create and load the alignment the alignment database. The gbrowse_syn_load_alignment_database.pl script is pre-installed with GBrowse.
$ cd .. $ mysql -uroot -pgbsyndemo -e 'create database orthocluster' $ gbrowse_syn_load_alignment_database.pl -u root -p gbsyndemo -d orthocluster -c -v orthocluster.txt
7) Copy the configuration files to the required location
$ cd conf $ sudo cp *conf /etc/gbrowse2/synteny
8) Go back to your browser and reload the rice page. There should now be a second data source in a pull-down menu.
9) Select the other data source and start browsing!
Further Reading
A Note on Whole Genome Alignments
The focus of the section of the course is on dealing with alignment or synteny data and using GBrowse_syn. However, how to generate whole genome alignments, identify orthologous regions, etc., are the subject of considerable interest, so some background reading is listed below:
- Primer on Hierarchical Genome Alignment Strategies
- article on PECAN and ENREDO
- all about PECAN
- The gene annotations for each species are in GFF files.
- The alignment data are in a constrained CLUSTALW format (They were not generated by the program CLUSTALW, which is not necessarily suitable for whole genome alignments)
- There are processing steps for the alignment data but it is very computationally intensive and we will load pre-processed data to get a head start.
Documentation
There is detailed documentation on the GMOD wiki for how to install, configure and use GBrowse_syn. To get started, browse these pages: