GBrowse syn Help
From GMOD
Contents
GBrowse_syn User Interface Help
GBrowse_syn is a GBrowse based synteny viewer. This page provides help on using GBrowse_syn. See the GBrowse_syn page for other information on GBrowse_syn.
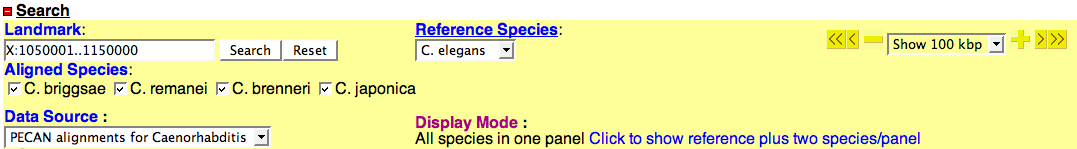
Search Section
Landmark
- The landmark input box accepts segment labels in the form:
reference sequence:start..end
- In some cases, gene names and other landmarks can also be entered. Support for searching other classes depends on the configuration for the species' data source.
- Note, make sure you have selected the correct reference species before clicking the 'Search' button.
Reference Species
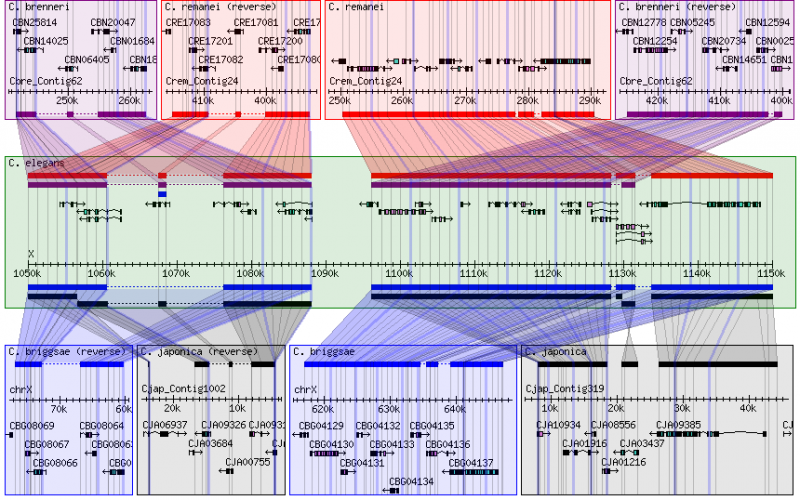
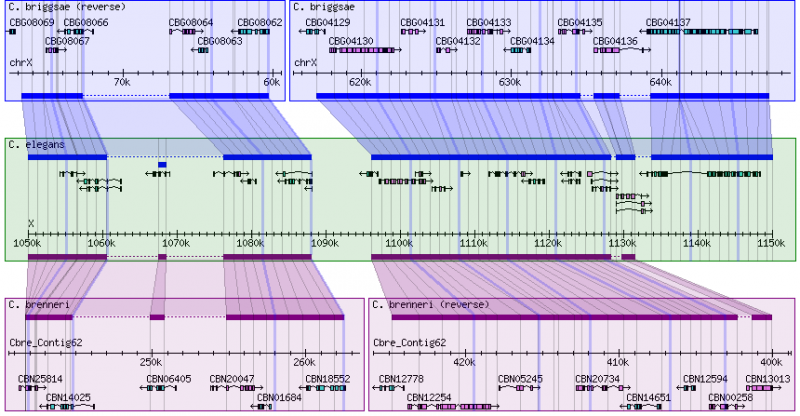
- This is the species that occupies the center panel in the alignment display.
- Alignments for other species are shown with reference to this coordinate system.
- Select the species from the pull-down menu and check boxes (see below) to select which species should be aligned to the reference sequence.
Aligned Species
- Configured species (except for the reference species) for the selected data source will be listed here.
- By checking each box, you indicate that alignments for this species, if available, should be displayed relative to the reference species.
Data Source
- This pull-down menu lists all available data sets configured for the synteny browser.
- Each item in this list corresponds to a sourcename.synconf configuration file.
- If only one data source is available this menu will not appear.
Display Mode
- The default display mode is to have the refernce species plus two aligned species per panel.
- This mode is best suited to displaying all sequences on roughly the same scale.
- The other display mode is
- The display mode can be toggled between expanded and compact by clicking the link shown above or via a pull-down menu on the "Display Settings" section