Difference between revisions of "GBrowse"
| Line 1: | Line 1: | ||
{{Tool data | {{Tool data | ||
| − | |||
|name=GBrowse | |name=GBrowse | ||
|full name=Generic Genome Browser | |full name=Generic Genome Browser | ||
| − | |status= | + | |status=mature |
| − | |dev= | + | |dev=inactive |
| − | |support= | + | |support=inactive |
|type=Genome Visualization & Editing | |type=Genome Visualization & Editing | ||
|platform=web | |platform=web | ||
| Line 27: | Line 26: | ||
|open source=Yes | |open source=Yes | ||
|licence=GPL2, Artistic License | |licence=GPL2, Artistic License | ||
| + | |input data type= | ||
|input=GFF3, GFF2 | |input=GFF3, GFF2 | ||
| + | |output= | ||
|language=Perl | |language=Perl | ||
| + | |audience= | ||
|release date=2001/01/01 | |release date=2001/01/01 | ||
| + | |use template=Yes | ||
|logo=GBrowseLogo.png | |logo=GBrowseLogo.png | ||
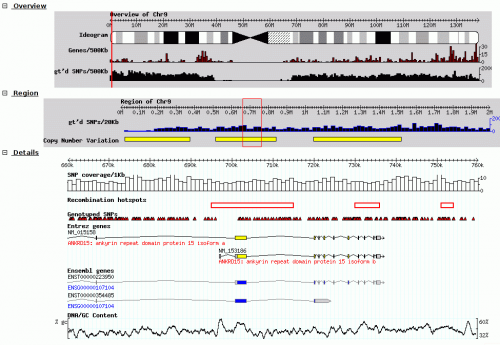
|screenshot=[[image:GBrowse_screenshot1.png|none|thumb|500px|GBrowse running on [http://hapmap.ncbi.nlm.nih.gov/downloads/index.html HapMap.org] [[Media:GBrowse_screenshot1.png|Click to view at full resolution]]]] | |screenshot=[[image:GBrowse_screenshot1.png|none|thumb|500px|GBrowse running on [http://hapmap.ncbi.nlm.nih.gov/downloads/index.html HapMap.org] [[Media:GBrowse_screenshot1.png|Click to view at full resolution]]]] | ||
| + | |contact= | ||
|mail={{MailingListsFor|GBrowse}}Please report bugs to the [http://sourceforge.net/tracker/?func=add&group_id=27707&atid=391291SourceForge Bug Tracker] (select 'Category: Gbrowse'). | |mail={{MailingListsFor|GBrowse}}Please report bugs to the [http://sourceforge.net/tracker/?func=add&group_id=27707&atid=391291SourceForge Bug Tracker] (select 'Category: Gbrowse'). | ||
| + | |dl= | ||
|dl src=GBrowse can also be found on [http://search.cpan.org CPAN] and in the Debian sid ("unstable") apt package repository and can be installed via apt-get in Ubuntu 12.04 and later. | |dl src=GBrowse can also be found on [http://search.cpan.org CPAN] and in the Debian sid ("unstable") apt package repository and can be installed via apt-get in Ubuntu 12.04 and later. | ||
|papers=* Using the Generic Genome Browser ([[GBrowse]]). <ref name=PMID:19957275/> | |papers=* Using the Generic Genome Browser ([[GBrowse]]). <ref name=PMID:19957275/> | ||
| Line 42: | Line 47: | ||
* Gbrowse Moby: a Web-based browser for [http://biomoby.open-bio.org/ BioMoby] Services. <ref name=PMID:17147784/> | * Gbrowse Moby: a Web-based browser for [http://biomoby.open-bio.org/ BioMoby] Services. <ref name=PMID:17147784/> | ||
* The [[GBrowse|generic genome browser (GBrowse)]]: a building block for a model organism system database. <ref name=PMID:12368253/> [[Media:1599-1.pdf|PDF]] | * The [[GBrowse|generic genome browser (GBrowse)]]: a building block for a model organism system database. <ref name=PMID:12368253/> [[Media:1599-1.pdf|PDF]] | ||
| − | + | |presentations= | |
|tutorials=; [[GBrowse Tutorial|GBrowse tutorial from 2012 GMOD Summer School]]. This relies heavily on the {{GBrowseAdminTutorialLink| GBrowse Administration Tutorial}}. | |tutorials=; [[GBrowse Tutorial|GBrowse tutorial from 2012 GMOD Summer School]]. This relies heavily on the {{GBrowseAdminTutorialLink| GBrowse Administration Tutorial}}. | ||
: Demonstrates setting up, configuring and using [[GBrowse]] with some sample data. GBrowse is provided on an Amazon Machine Image; see [[Cloud|GMOD in the Cloud]] for more information on getting a GMOD AMI. | : Demonstrates setting up, configuring and using [[GBrowse]] with some sample data. GBrowse is provided on an Amazon Machine Image; see [[Cloud|GMOD in the Cloud]] for more information on getting a GMOD AMI. | ||
| Line 57: | Line 62: | ||
; [http://www.openhelix.com/gbrowse GBrowse User Tutorial] at [http://www.openhelix.com OpenHelix] | ; [http://www.openhelix.com/gbrowse GBrowse User Tutorial] at [http://www.openhelix.com OpenHelix] | ||
: A Flash based tutorial on using GBrowse. Provided by [http://www.openhelix.com OpenHelix]. Tutorial includes slides, handouts and exercises. NB: this tutorial is for [[GBrowse 1.x]] | : A Flash based tutorial on using GBrowse. Provided by [http://www.openhelix.com OpenHelix]. Tutorial includes slides, handouts and exercises. NB: this tutorial is for [[GBrowse 1.x]] | ||
| + | |getting started preamble= | ||
|req=GBrowse is [[Glossary#Perl|Perl]]-based and the GBrowse 2.x modules are [http://search.cpan.org/dist/GBrowse/ hosted on CPAN]. GBrowse can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed. | |req=GBrowse is [[Glossary#Perl|Perl]]-based and the GBrowse 2.x modules are [http://search.cpan.org/dist/GBrowse/ hosted on CPAN]. GBrowse can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed. | ||
|install=* [[GBrowse_2.0_Install_HOWTO|GBrowse 2.x installation guide]] | |install=* [[GBrowse_2.0_Install_HOWTO|GBrowse 2.x installation guide]] | ||
| Line 159: | Line 165: | ||
* [http://search.cpan.org/dist/GBrowse/docs/pod/GBROWSE_IMG.pod GBROWSE_IMG] | * [http://search.cpan.org/dist/GBrowse/docs/pod/GBROWSE_IMG.pod GBROWSE_IMG] | ||
* [http://search.cpan.org/dist/GBrowse/docs/pod/ORACLE_AND_POSTGRESQL.pod ORACLE_AND_POSTGRESQL] | * [http://search.cpan.org/dist/GBrowse/docs/pod/ORACLE_AND_POSTGRESQL.pod ORACLE_AND_POSTGRESQL] | ||
| + | |dev ppl= | ||
| + | |dev ppl= | ||
| + | |dl tracking= | ||
|logo info=The [[:File:GBrowseLogo.png|GBrowse logo]] was created by [mailto:alexisnb1@yahoo.com Alex Read], a participant in the [[Spring 2010 Logo Program]], while a design student at [http://www.linnbenton.edu Linn-Benton Community College]. | |logo info=The [[:File:GBrowseLogo.png|GBrowse logo]] was created by [mailto:alexisnb1@yahoo.com Alex Read], a participant in the [[Spring 2010 Logo Program]], while a design student at [http://www.linnbenton.edu Linn-Benton Community College]. | ||
|see also=[[JBrowse]], the successor to GBrowse, built with JavaScript for a faster, more interactive user experience; and [[WebGBrowse]], a tool for configuring GBrowse. | |see also=[[JBrowse]], the successor to GBrowse, built with JavaScript for a faster, more interactive user experience; and [[WebGBrowse]], a tool for configuring GBrowse. | ||
| Line 167: | Line 176: | ||
|dev status= | |dev status= | ||
}} | }} | ||
| − | + | [[Category:GMOD_Components]] | |
| + | [[Category:GBrowse]] | ||
{{SemanticLink | {{SemanticLink | ||
|linkurl=http://sourceforge.net/projects/gmod/files/Generic%20Genome%20Browser/ | |linkurl=http://sourceforge.net/projects/gmod/files/Generic%20Genome%20Browser/ | ||
| + | |linktitle= | ||
|linktype=download | |linktype=download | ||
| + | |linkdesc= | ||
}} | }} | ||
{{SemanticLink | {{SemanticLink | ||
|linkurl=https://github.com/GMOD/GBrowse | |linkurl=https://github.com/GMOD/GBrowse | ||
| + | |linktitle= | ||
|linktype=source code | |linktype=source code | ||
| + | |linkdesc= | ||
}} | }} | ||
{{SemanticLink | {{SemanticLink | ||
|linkurl=http://gbrowse.org | |linkurl=http://gbrowse.org | ||
| + | |linktitle= | ||
|linktype=website | |linktype=website | ||
| + | |linkdesc= | ||
}} | }} | ||
{{SemanticLink | {{SemanticLink | ||
| Line 184: | Line 200: | ||
|linktitle=WormBase | |linktitle=WormBase | ||
|linktype=wild URL | |linktype=wild URL | ||
| + | |linkdesc= | ||
}} | }} | ||
{{SemanticLink | {{SemanticLink | ||
| Line 189: | Line 206: | ||
|linktitle=FlyBase | |linktitle=FlyBase | ||
|linktype=wild URL | |linktype=wild URL | ||
| + | |linkdesc= | ||
}} | }} | ||
{{SemanticLink | {{SemanticLink | ||
| Line 194: | Line 212: | ||
|linktitle=HapMap | |linktitle=HapMap | ||
|linktype=wild URL | |linktype=wild URL | ||
| + | |linkdesc= | ||
}} | }} | ||
| − | |||
| − | |||
| − | |||
Latest revision as of 16:47, 10 April 2023
- Mature release
- Development: inactive
- Support: inactive
Contents
About Generic Genome Browser (GBrowse)
GBrowse is a combination of database and interactive web pages for manipulating and displaying annotations on genomes. Features include:
- Simultaneous bird's eye and detailed views of the genome.
- Scroll, zoom, center.
- Use a variety of premade glyphs or create your own.
- Attach arbitrary URLs to any annotation.
- Order and appearance of tracks are customizable by administrator and end-user.
- Search by annotation ID, name, or comment.
- Supports third party annotation using GFF formats.
- Settings persist across sessions.
- DNA and GFF dumps.
- Connectivity to different databases, including BioSQL and Chado.
- Multi-language support.
- Third-party feature loading.
- Customizable plug-in architecture (e.g. run BLAST, dump & import many formats, find oligonucleotides, design primers, create restriction maps, edit features)
Note that the information on this page refers to GBrowse 2; GBrowse 1.x is recommended only for applications where legacy browser support is required and a single database is used.
Visit the GBrowse website.
Screenshots
Downloads
GBrowse can also be found on CPAN and in the Debian sid ("unstable") apt package repository and can be installed via apt-get in Ubuntu 12.04 and later.
Using GBrowse
System Requirements
GBrowse is Perl-based and the GBrowse 2.x modules are hosted on CPAN. GBrowse can be installed using the standard Perl module build procedure, or automated using a network-based install script. In order to use the net installer, you will need to have Perl 5.8.6 or higher and the Apache web server installed.
Installation
Configuration
Documentation
- GBrowse FAQ
- Annotation Help
- Balloons:
- GBrowse Persistent Variables
- GBrowse img
- Glyphs and Glyph Options
- Rubber Band Selection
- Gbrowse Benchmarking
- GBrowse User Uploads
POD documentation
There are many useful POD documents included with the distribution. These are converted to HTML files when you install the package, and can be found in /gbrowse/docs/pod.
GBrowse 2.x pod documents can also be viewed online at CPAN:
- FAQ
- INSTALL
- INSTALL.MacOSX
- README-chado
- README-gff-files (see also GFF)
- README-lucegene
- BIOSQL_ADAPTER_HOWTO
- GENBANK_HOWTO
- PLUGINS_HOWTO
- DAS_HOWTO
- MAKE_IMAGES_HOWTO
- GBROWSE_IMG
- ORACLE_AND_POSTGRESQL
Installation
Configuration
Documentation
- GBrowse FAQ
- Annotation Help
- Balloons:
- GBrowse Persistent Variables
- GBrowse img
- Glyphs and Glyph Options
- Rubber Band Selection
- Gbrowse Benchmarking
- GBrowse User Uploads
POD documentation
There are many useful POD documents included with the distribution. These are converted to HTML files when you install the package, and can be found in /gbrowse/docs/pod.
GBrowse 2.x pod documents can also be viewed online at CPAN:
- FAQ
- INSTALL
- INSTALL.MacOSX
- README-chado
- README-gff-files (see also GFF)
- README-lucegene
- BIOSQL_ADAPTER_HOWTO
- GENBANK_HOWTO
- PLUGINS_HOWTO
- DAS_HOWTO
- MAKE_IMAGES_HOWTO
- GBROWSE_IMG
- ORACLE_AND_POSTGRESQL
Publications, Tutorials, and Presentations
Publications on or mentioning GBrowse
- Using the Generic Genome Browser (GBrowse). [1]
- SNP@Evolution: a hierarchical database of positive selection on the human genome. [2] (GBrowse-related)
- TBrowse: an integrative genomics map of Mycobacterium tuberculosis. [3] (GBrowse-related)
- FishMap: a community resource for zebrafish genomics. [4] (GBrowse-related)
- Using the Generic Genome Browser (GBrowse). [5]
- Genome-Wide Analysis of Nucleotide-Level Variation in Commonly Used Saccharomyces cerevisiae Strains. [6] (See YSB.)
- Gbrowse Moby: a Web-based browser for BioMoby Services. [7]
- The generic genome browser (GBrowse): a building block for a model organism system database. [8] PDF
Tutorials
- GBrowse tutorial from 2012 GMOD Summer School. This relies heavily on the GBrowse2 Admin Tutorial.
- Demonstrates setting up, configuring and using GBrowse with some sample data. GBrowse is provided on an Amazon Machine Image; see GMOD in the Cloud for more information on getting a GMOD AMI.
- GBrowse tutorial from 2010 GMOD Summer School
- Set up and run GBrowse with sample data. It provides a VMware image to work on, and relies heavily on the GBrowse2 Admin Tutorial.
- GBrowse2 Admin Tutorial
- Step by step guide on how to configure and load data into GBrowse. Administration tutorials are available for both the GBrowse2 Admin Tutorial, and the earlier 1.x versions.
- Usage tutorial
- GBrowse usage tutorial
- GBrowse NGS Tutorial
- Instructions on how to visualize next generation sequencing data in GBrowse using SAMtools. The tutorial includes a starting VMware image, and uses the example data that comes with SAMtools.
- GBrowse video tutorial
- Produced by EuPathDB; please direct praise and thanks to them!
- GBrowse User Tutorial at OpenHelix
- A Flash based tutorial on using GBrowse. Provided by OpenHelix. Tutorial includes slides, handouts and exercises. NB: this tutorial is for GBrowse 1.x
Contacts and Mailing Lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| GBrowse & GBrowse_syn | gmod-gbrowse | GBrowse and GBrowse_syn users and developers. | Gmane, Nabble (2010/05+), Sourceforge |
| gmod-gbrowse-cmts | Code updates. | Sourceforge |
GBrowse in the wild
Public installations of GBrowse:
See also
JBrowse, the successor to GBrowse, built with JavaScript for a faster, more interactive user experience; and WebGBrowse, a tool for configuring GBrowse.
More on GBrowse
See Category:GBrowse
GBrowse Logo
The GBrowse logo was created by Alex Read, a participant in the Spring 2010 Logo Program, while a design student at Linn-Benton Community College.
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:19957275 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:19732458 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:19683474 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:18554176 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:18428797 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:17389913 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:17147784 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedPMID:12368253
| Available on platform | web + |
| Has URL | http://sourceforge.net/projects/gmod/files/Generic%20Genome%20Browser/ +, https://github.com/GMOD/GBrowse +, http://gbrowse.org +, http://www.wormbase.org/tools/genome/gbrowse/c_elegans/ +, http://flybase.org/cgi-bin/gbrowse/dmel + and http://hapmap.ncbi.nlm.nih.gov/cgi-perl/gbrowse/gbrowse + |
| Has description | GBrowse is a combination of database and i … GBrowse is a combination of database and interactive web pages for manipulating and displaying annotations on genomes. Features include:
|
| Has development status | inactive + |
| Has full name | Generic Genome Browser + |
| Has input format | GFF3 + and GFF2 + |
| Has licence | GPL2 + and Artistic License + |
| Has logo | GBrowseLogo.png + |
| Has software maturity status | mature + |
| Has support status | inactive + |
| Has title | WormBase +, FlyBase + and HapMap + |
| Has topic | GBrowse + |
| Is open source | Yes + |
| Link type | download +, source code +, website + and wild URL + |
| Release date | 1 January 2001 + |
| Tool functionality or classification | Genome Visualization & Editing + |
| Written in language | Perl + |
| Has subobjectThis property is a special property in this wiki. | GBrowse#http://sourceforge.net/projects/gmod/files/Generic%20Genome%20Browser/ +, GBrowse#https://github.com/GMOD/GBrowse +, GBrowse#http://gbrowse.org +, GBrowse#http://www.wormbase.org/tools/genome/gbrowse/c_elegans/ +, GBrowse#http://flybase.org/cgi-bin/gbrowse/dmel + and GBrowse#http://hapmap.ncbi.nlm.nih.gov/cgi-perl/gbrowse/gbrowse + |